1 Introduction
As the application of Next Generation Sequencing (NGS) technologies contributes to the generation of ever larger amounts of new data, to take full advantage of all this new knowledge we need to build structures to organize, process and use it in order to improve our understanding of the human genome.
Previous studies [1-3] have shown how conceptual models allow us to provide a definition of a domain, so that we can understand the entities involved and their relationship. These studies focused on describing the genome of eukaryotic cells and the interaction between proteins, transcriptome, and other genetic components. Other studies focus specially on proteins by Ram [23]. Pastor et. al. describes the Conceptual Schema of the Human Genome (CSHG) [4], [24]. However, this conceptual schema requires to be constantly aligned with the new genomic knowledge and in this paper we extend the aforementioned model to include the specification of haplotypes (which are defined in Section 2). For haplotypes, this model should be extended in two ways: 1) integrating treatment of haplotypes, 2) application of statistical models.
In this context, the goal of the present study, which is based on our previous work [48], is to extend the Conceptual Schema of the Human Genome (CSHG) by including the concepts of haplotypes and statistical models, thus improving the schema’s expressiveness. This way, we foresee the creation of a powerful and reliable Genomic Information System based on this holistic conceptual schema (CS).
The advances over our previous work [48] are:
The description of the preliminary steps achieved in the treatment of haplotype data: e.g., the study of repositories, data collection and analysis, data loading and some test queries, and
The explanation of how the different representations of genomic knowledge affect the structure of the underlying database that is used to manage all the data. This scenario shows how essential it is to have a sound understanding of the relevant information to achieve efficient data management policies.
This paper is divided as follows: Section 2 gives the background to this work and defines the concept of haplotypes. Section 3 explains the research methodology used. Section 4 reviews related work on how haplotype information is currently structured and stored in leading genomic data repositories. Section 5 contains our proposal for the conceptual modeling of haplotypes. Section 6 describes the conceptual alignment that we suggest to formalize haplotype information. Section 7 describes the first steps in haplotype data treatment. Section 8 contains a discussion on database evolution according to the Conceptual Schema of the Human Genome (CSHG). Finally, section 9 presents the lessons learned and outlines future work.
2 Background: Understanding the Haplotype Concept -test case: Alcohol Sensitivity-
We detected the importance of including haplotype treatment in our CSHG on the genetic implications for the pathology of Alcohol Sensitivity, in which we did an intensive study of genes and variants that were associated with a predisposition to this disease [5-6].
Alcohol sensitivity occurs when an individual ingests a certain amount of alcohol causing immediate rejection and experiencing discomfort, dizziness and other symptoms. The simple fact of consuming thus causes discomfort and has a future impact on the health of the individual [11], [58]. We have taken this disease as a test case because it occurs in the population as a whole, regardless of social status, age or culture [8].
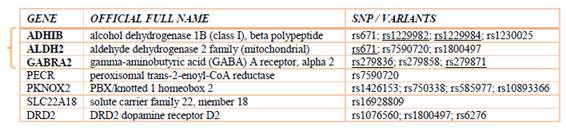
For the study of these genetic alterations we obtained a lot of information from different repositories of genomic data (such as NCBI: dbSNP [57], PubMed [59], and others). We defined a group of genes [6] closely linked to the disease, including:
These genes are directly or indirectly associated with alcohol sensitivity. The genes with a direct influence present a high the predisposition to this disease, and the indirect are in a more general perspective (because some are associated with addictive diseases).
After completing the search and identification processes [38], we turned to the medical validation of genes, with the help of our fellow biologists (supporting this validation with papers from high-impact medical journals). We proceed to filter the genes in Table 1, selecting only three relevant genes: (a) ALDH2 (rs671); (B) GABRA2 (rs279836, rs279871) and (c) ADH1B (rs1229982, rs1229984).
We detected a "haplotype case" with the GABRA2 gene, in which we found a haplotype composed of three variants (variations): rs279871, rs279836 and rs279845 [38]. Initially we worked with individual variations , without considering the relationships between them and without considering the rs279845 variant [11-12]).
Variation (or variants): naturally occurring genetic differences among organisms in the same species [Scitable by Nature Edu.].
From a biological point of view, Haplotypes are a set of SNPs that are inherited and found together in a chromosome and are defined as a group of SNPs of a gene that are very close and tend to be inherited together. This means that a haplotype of the alleles is not separated in the recombination phase and can be transmitted in blocks, allowing combinations of variants to a gene that affects certain phenotypes [7].
There is currently a significant set of genetic diseases in which the influence of haplotypes has been well established, such as breast cancer [9-10] and alcohol sensitivity [11-12], among others [13-14]. If haplotypes are considered in the diagnosis, the outcome can be improved, as the assessed probabilities indicate the level of risk in genomic diagnosis more accurately.
After making this discovery we realized that our conceptual schema should be improved to achieve an integration of information in a more appropriate manner. If we treat this concept effectively, we can improve the results generated in the genetic diagnoses.
Figure 1 shows the differences in genetic analysis focused only on individual variants against studies on variants plus haplotypes in a reference sample. As the figure explains (in Path A) in this situation the doctor looks for a change in the sample associated with a specific disease; but in fact merely checks if the variation (SNP) exists in the analyzed sample. With this information a report is generated on the existence of the variation.
Path B in the figure explains the ideal process we address in this work. The doctor looks for the variants and determines whether there are combinations between all the variants, trying to find haplotypes on different alleles through the analysis of the frequencies of each one. In order to obtain this data, we should apply different statistical models to present a more detailed and comprehensive genetic report. It is widely accepted that Haplotype studies enhance the variation detection rate (with or without combinations) for a specific disease [17]. The reason is that each allele or variation represents a frequency of occurrence in each population, so that with this information we can improve the generation of genetic diagnosis.
2 A single nucleotide polymorphism (SNP), is a variation at a single position in a DNA sequence among individuals. It should be remembered that the DNA sequence is formed from a chain of four nucleotide bases: A, C, G, and T [Scitable by Nature Edu.].
3 Research MethodologyFigure 2
This paper deals with and analyzes only a part of a wider PhD research project and is based on the presentation of “Treatment Design” [15]. Accordingly, we used the Engineering Cycle method proposed by Wieringa [16], which includes the following steps:
a) Implementation evaluation / Problem investigation
In this phase we focus on describing the problem and submit initial proposals for the resolution of the case [15]. Our work with haplotypes and variants explains the necessity of extending our CSHG, because all the possible combinations between variants will help us to improve the impact of the diagnosis of genetic diseases.
b) Treatment design
One of the steps taken to design our solution was to study in depth the state of the art of the subject and then proceed to apply analysis and evaluation techniques to obtain the relevance and the impact factor of haplotypes for genetic diagnosis. Our solution is based on an integrative approach to heterogeneous genomic data sources using conceptual models as the main formalism. Our goal is to gather and represent the available knowledge using an unambiguous representation and establish concise relationships among the data. From these models we will develop a GeIS using a Model-driven development (MDD) approach [41].
c) Treatment validation
Design validation is a knowledge task [15]. We are going to provide validation mechanisms for the outcome from our CSHG (which integrates the haplotype treatment into the genomic diagnosis) developed in the previous step.
d) Treatment implementation
Finally, the implementation of our solution will be tested by a selected group of physicians working in the genetic analysis domain and also with a group of users to obtain feedback on the functionality and the value of the information provided.
In the context of this paper we give an overview of the research and development of the second step of the Engineering Cycle (Treatment Design), in which we address the following points (research phases):
1. Study and analysis of the state of the art (Section 4)
2. Study of the domain and analysis of relevant information (Sections 2 and 4)
3. Extension of the conceptual schema and the design of Information Systems (Section 5)
4. Integration of relevant information into a single genomic database (Sections 6 and 7)
4 Related WorkTable 2
Little work has been done to date on modeling "haplotypes". Various studies have tried to achieve a conceptual definition of the entire human genome, but this knowledge quickly becomes obsolete due to the continuous evolution of the domain.
Our research focuses on the integration and use of information on haplotypes in genomic information repositories, such as information systems and databases. For these reasons, we first analyze some of the most important genomic repositories in order to evaluate their schemas, the concepts they use and how haplotypes are stored.
dbSNP
The dbSNP repository is the top source of information on SNPs. This repository facilitates an ER schema which identifies the representation of data on: population, SNP allele frequencies and the summary of populations. dbSNP collects this data in the view "Frequency calculation submitted by SNP and population" with construction number 118 dated 11/17/2003. Within this view we find a table called "b125_SNPMapInfo_35_1" related to the "SNP" table, where there is only one attribute related with haplotypes - “hap_cnt”- [45], [56-57].
Ensembl
The Ensembl repository mainly provides genomes for vertebrate species. In this repository the schema does not provide any explicit relationship with haplotypes, but it is noteworthy that in the "features_analyses_core" view we detected some entities that could be linked to treatment, such as: the table "Marker_map_location" and the attribute "lod_score"; which is statistical data used in population genetics and in LD calculations [36], [46].
UCSC Genome Bioinformatics
This site contains reference sequences and working draft assemblies for a large collection of genomes. In this case, they present the data as a “Table schema” that is somewhat difficult to manage [54]. Using the "Gene Sorter" tool [40] we could check the different data provided for a gene, including data associated with haplotypes in the block "Common Gene haplotype Alleles", which are generated from the 1000 Genomes project (http://www.1000genomes.org/). From the data presented we drafted an outline of the structure to compare with our solution [39], [47].
We also found a set of databases focused on collecting data associated with haplotypes and population frequencies, such as: HapMap [18], ALFRED [19], YHRD [20], D-HaploDB [21], and others [4], [8]. The problem with this type of repository is that it is difficult to manage and access the haplotype information, as this information is widely dispersed, e.g., the data is stored in multiple text files (* .txt, * .csv, etc.). After analyzing different data sources (and schemas) that store haplotype information, we identified three main issues:
1. Complexity of data management:
Data is presented ambiguously, and in many cases is difficult to understand and manipulate, e.g., in the case of dbSNP and Ensembl, the haplotype data is not shown explicitly to final users. In our study we detected that the dbSNP repository uses data from the HapMap project, whereas USCS uses data from the 1000 Genomes project.
We also found some inconsistencies between these repositories (i.e., contradictory or inconsistent information in the databases, like allele and genotype frequencies). As the genomic environment is continuously evolving, additional knowledge is incorporated. Several sources show data with information on haplotypes, as in the cases mentioned above, but the problem lies primarily in the complexity of management and interpretation of the data (importance, relevance, etc.) [43].
2. High dispersion and data redundancy:
This issue is a consequence of different data sources with large amounts of structured and unstructured information in different formats, for example, formats like: *.csv; *.txt; *.xml; *.fasta; and others. This wide range of formats makes it very difficult to process and analyze the data, so it is reasonable to adopt in this domain the benefits of conceptual schemas, which allow us to create a structure in which data can be shared effectively and redundancy and other issues can be reduced [52].
A further disadvantage identified in the haplotype data is its wide dispersion and the presence of redundant data [43]. Using a conceptual modeling approach, we will tackle these problems with comprehensive data processing to complement existing genetic diagnostics.
3. There is no clear formalization of the concept "haplotypes":
Currently, the analyzed repositories do not provide a suitable structure (schema) to manage haplotypes. In some cases, they do not even represent the same concept, e.g. we found a lot differences between dbSNP and Ensembl on how they represent haplotypes at the conceptual level (i.e. form of representation, structure and others). We only found a sort of table schema specification in UCSC. dbSNP only shows an attribute associated with the concept of haplotypes in its schema, and for this reason is a very limited definition. Ensembl does not provide a clear definition in its schema. We found the "lod_score" attribute that is used in genetics, but not specifically for haplotype treatment. Although UCSC presents data on haplotypes, this repository does not have a conceptual schema.
There are also other alternatives for representing knowledge in general, and in our research we found ontologies applied to biological sequences. Sequence Ontology is a set of terms and relationships used to describe the features and attributes of biological sequences (http://www.sequenceontology.org/). This ontology defines a haplotype as one of a set of coexisting sequence variants of a haplotype block [55] and this approach is interesting for defining types, properties and relationships between entities at a more formal specification level. Our solution seeks to: (i) represent the existing data in this domain, manipulate and manage the information on haplotypes so as to make them easy to use in genomic treatment, (ii) solve existing shortcomings in this domain through the practical application of conceptual schemas, which may be open to extension regardless of the continuing evolution of genomic environment.
5 Conceptual Modeling of Haplotypes
The application of data management techniques in a genomic environment could be thwarted or affected by its special characteristics, such as: high conceptual complexity, large amounts of data and the constant evolution of the community.
Our research group has been able to generate a Conceptual Schema of the Human Genome (CSHG) [24] which has evolved and grown over the years and has advanced the understanding of the human genome.
The latest version of the CSHG (v3) is classified into five views [44], which are:
(1) Structural view: this describes the genome structure (species, chromosomes, etc.).
(2) Transcription view: shows the components and concepts related to protein synthesis. Components involved in going from DNA to the diversity of RNAs.
(3) Variation view: this view models the knowledge related to the differences in the DNA sequence of different individuals.
(4) Pathway view: there are a series of chemical reactions within a cell in the area of biochemical pathways or metabolic pathways intended to enrich the conceptual schema with information on metabolic pathways joining genome components that participate in pathways with phenotype expressions.
(5) Bibliography and data bank view: gives the source of the data and contains a series of bibliographic reference documents for anyone wishing to obtain further information [22], [24].
Our idea of integrating haplotypes strengthens the conceptual schema and also assesses levels of incidence or the risk of "variations" in predisposing genetic diseases. Figure 3 shows the current state of the “variation view”, indicating classes (grey boxes) that are currently loaded into our repository (using a data loading process [42], [49]).
In our CSHG (v3) we now use precise variations (i.e. when the structure and the nucleotides that are involved are clearly defined). However, when treating haplotypes, other concepts must also be managed, including: frequencies (allelic and genotypic) and populations [25], which is difficult to manage in the genomic field. This new scenario means that the data loading process should also consider the concepts represented on the left-hand side (Frequency generalization) of Figure 3.
The variations are presented in two groups divided by: frequency and description. The first is classified according to the variation frequency in the population (occurrence). In this case, there are two types: mutations and polymorphisms. An alteration in any part of the genetic code is known as a mutation and a single nucleotide change between the genomes of individuals of the same species is known as polymorphism (occurring in at least 1% of the population). The second group conforms to the description given by the variations, classified into two types: precise and imprecise. The former consists of those that indicate their position in the chromosome and the latter only describe their phenotypic effect but do not give their location within the chromosome.
The "SNP" class of our CSHG becomes the root class where new concepts appear. Although the conceptualization of "SNP_Allele"; "SNP_Allele-Pop"; "SNP_Genotype"; "SNP_Genotype-Pop"; "Population"; "LD" classes is represented in our original proposal, the data related to them was neither treated nor loaded into our repository for one of two causes: (a) unavailability of the data (sources, resources, etc.) or (b) we did not consider it to have an appreciable or tangible value.
An SNP is associated with many alleles ("SNP Allele") and these sets have a frequency of alleles in a specific population ("SNP_Allele-Pop"). Similarly, it also has several types of genotypic data ("SNP_Genotype") defined by "allele1" and "allele2" (the reference allele and the allele changed within the genotype); each allele represents a frequency within a population. The "Population" class is used to group all the populations that have been studied for the analysis of variants (variations) in the human genome.
Figure 4 gives our proposal for integrating haplotypes into the CSHG, in which we found a number of added or extended classes. As can be seen, the variation view defined by their "description" is preserved, as in the current version. In this figure, variations defined by “frequency” have to implement changes that are necessary for working with the haplotype data and statistical models.
The insertions and changes made to the schema are explained as follows: the "Haplotype" class, which is associated with the "SNP" class, is added, reflecting the relationships present in a Haplotype-SNP by the combination of two or more SNPs. The attributes that define "Haplotype" class in the schema are: "id_reference" which defines an identifier a link between the different SNPs and haplotype; the "id_haplotype" corresponds to the identifier of the haplotype as the "variation_id" that identifies the associated variation. As the haplotype should be based on a scientific resource to confirm its medical value, we created the class called "Hap_Bib_Ref" to help us the join the “Haplotype” and "Bibliography reference" classes, to keep our repository linked to various research works on haplotypes.
This class consists of the following attributes: "population_id" that serves as an identifier of the population; the identifier of the source "pop_db_id"; the name of the population "pop_name" and geographical region "geo_region". We also associated the new "Pop_OriginDB" class, which we use to define the sources that provide the above populations. In this class we define the following attributes: "pop_orig_id" which is the identifier of the repository; the "name" and "description" of the source, and the "URL_pop" which has the URL of population data file.
Another novelty in the schema is the addition of the "Statistical_models" class, which was defined with the objective of unifying the statistical models that are applied to the data related with variations [26-28], specifically on changes in the value of "variation_id", considering that for an SNP or variation this can be zero or many statistical models. Three subclasses of this superclass are derived: "SNP_HardyW", "Fst" and "LD".
The conceptual definition of these classes allows us to tackle very important concepts in the world of genomics, known as "Population Genetics", which is the study of the forces that alter the genetic makeup of a species. This focus is associated with micro-evolutionary mechanisms like: mutation, natural selection, gene flow and genetic drift [29-30]. The "SNP_HardyW" class represents the Hardy-Weinberg model (also known as panmictic balance), which is used to calculate genotype frequencies from allele frequencies [31-32], in which data is taken from the sources and applied to the model. This class has the following characteristics: unique identifier of the "pbal_id" class; chromosomal position in base pairs "pos_bp"; and the P-value "pvalue" which indicates the smallest possible level of significance.
Hardy-Weinberg equation3 = p2 + 2pq + q2 = 1
Where: p: is the frequency of the "A" allele and q is the frequency of the "a" allele in the population. In the equation,
p2: frequency of the homozygous genotype AA,
q2: frequency of the homozygous genotype aa, and
2pq: frequency of the heterozygous genotype Aa.
Another statistical value used in population genetics is the calculation of "fixation indices", which allow us to measure the differentiation of the population due to genetic structure, facilitating the comparison of genetic variability within and between populations. To do this we define the "Fst" class, containing the values: "Fst" for fixation index and the "Ave_Het", which indicates the average heterozygosity. The "LD" class defines the "Linkage disequilibrium", which occurs when the genotypes in the two loci are not independent of each other.
To calculate the LD, we found three statistical biological parameters, which are: (1) the sensitivity index "DPrime", which measures the imbalance between the alleles’ interaction. (2) The coefficient of determination "RSquare" which serves to determine the quality of the model, in order to replicate the results, and the proportion of variation in the results presented in the model. (3) LOD score "LOD": this value refers to the logarithm on the odds of two genes or loci being linked and thus being inherited together more often than usual.
The "LD" class is related to the "SNP" class, indicating that an SNP can have zero to many LDs. In this extension of the CSHG we integrated all the existing data on phenotypes from different repositories and suggested the "Phenotype" class, which is joined to the "Variation" class through the intermediate class "Certainty", used as the indicator of the incidence level between phenotype-variation (this value is extremely difficult to define, but studies on this topic provide "estimated values" within the population).
Finally, the "Gen_non_redund" class helps us to offer the results of the cleaned dataset (without redundancy, i.e. elimination of inconsistencies and duplicated data), for sets for SNP-genotyping and population. For this class we assign a unique identifier to the data without redundancy "gen_nonred_id"; information about chromosome and the studied population, the attributes are "chromosome" and "population_id" respectively; and total non-redundant data after performing filtering "gen_non_redund".
3 Hardy-Weinberg equation: this is a mathematical equation that can be used to calculate the genetic variation of a population at equilibrium [Scitable by Nature Education].
6 Conceptual model validation
To validate our work, we checked whether the extended conceptual model supports the information provided by popular haplotypes repositories: establishing conceptual alignment with data available from popular genomic repositories. Figure 4 shows the elements added to the schema and their sources. The main data source was the HapMap Project in its third phase.
According to our variation view, the HapMap dataset is contained in the following directories: (a) frequencies and (b) ld_data. First, “frequencies” is a directory that contains the above SNPs with their frequency in each population. Due to the large amount of information, it is clustered into two groups: the frequencies of SNP variant types, taking into account only one allele of the chromosome ("Allele_freqs"), and frequencies of SNP rate variants, taking into account the two alleles ("Genotype_freqs").
The table below shows the elements added to the schema, the data repositories and the specific field/table which is aligned with the new element.
The Linkage disequilibrium data [33] is provided by the "LD_data" directory. The large amount of information is divided by chromosomes and populations. Information on population "Population" and non-redundant genes "Gen_non_redund" is found in specific sections of the main website.
The "Statistical_models" class consists of the data extracted from HapMap for "LD", and provided by the ALFRED repository for frequencies and statistical processes applied to the biological field. The frequencies dataset from ALFRED is in a file called "FreqByChrom" available from their website; these frequencies have been obtained for each chromosome according to the population studied.
This repository also facilitates the extraction of information on the populations they used. For the “phenotypic” entity, we find several data sources from which we can extract all the information classified in multiple repositories, such as: ClinVar, Ensembl and DisGeNET (sorted by their scientific relevance) [34-37].
7 Development of a haplotype databaseFigure 8
This section deals with the treatment of data (haplotypes) from the previously defined conceptual model (Figure 4), using the data repositories to define queries relevant to the analysis of the data. As mentioned in the previous section, the data on variations that are part of one or more haplotypes and their respective frequencies were obtained from different repositories. Our data management approach follows five steps:
a) Impact study of existing and available repositories
In this activity we focus on evaluating the usefulness of the data provided and its impact on research and scientific publications, with the aim of defining a framework data source according to the latest advances in the genomic context (see datasources in Table 3).
b) Data collection (haplotype frequencies and statistical calculations)
After selecting the data repositories, we proceed to download the files (specified in Table 3). At this stage it is usual to find huge data files, for example:
Allele frequencies for chromosome 1 with the studied HapMap populations (11) with an average of 450-1,024MB ±.
LD data associated with chromosome 1 for the "ASW" population with an average of 930-1,024MB ±.
In the case of ALFRED repository, we obtained the chromosomes frequencies with an average of 500-650MB ±. For files with phenotypic data it was much more affordable in size issues.
After downloading all the files, we generate multiple gigas of scattered information with completely different structures to be evaluated. In order to strengthen previous analyses, we decided to complement and compare the information with data generated from the BioQ platform (http://bioq.saclab.net/) [61], which provides a set of tools to consult, document and download information from relational databases (genomic), such as: 1000 Genomes, dbSNP, Ensembl, and others (for specific versions that they manage).
c) Analysis of the data stored
Due to the heterogeneity of the data, the first thing we did in this phase was to transform all the data from its current format (see Section 4) to a common structure. We decided to convert all the files to "*.csv" format for debugging and analysis, and then took the first genome chromosomes (i.e., 1-3) as a test case for handling and storage issues.
As we can see in Figures 7 and 8, after concluding our analysis of the relevant data for the treatment of haplotypes and statistical factors, we identified the large contribution of the ALFRED (1,063.651 rows) and HapMap (194.417 rows) repositories to our study. We also classified all the knowledge obtained into five categories (frequencies, statistical values, phenotypes, populations and others). The largest amount of the processed data was on frequencies and statistical values.
d) Massive data load
After completing the analysis and treatment of the data, we did a preliminary data filtering, and the next step was to develop an initial database schema for loading all the information. For this, we studied the most appropriate data structures to organize, consult and avoid processing problems (e.g. special signs) in the data.
The next task was to import the files (*.csv) using a database management environment (see Figure 9), and then generated a new ER diagram, while maintaining the traceability of the origin of the data used (see Figure 6).
e) Query generation (preliminary)
In this phase we raised a number of questions with the aim of finding answers through the generation of different SQL queries on the previously loaded schema, including the following examples:
(1)SELECT COUNT(DISTINCT(id_symbol)) FROM phenotype
In this query we included a total of 4,192 genes loaded, which represent an association with one or more diseases or phenotypes.
(2)SELECT phenotype_id, id_symbol, disease_name_descr, type, score, num_of_pubmed, source_name, diseaseMIM, last_updated, notes FROM phenotype WHERE id_symbol = 'GABRA2'
With this query we obtained the GABRA2 gene with the identifier 72.387 (phenotype_id), which was loaded from the ClinVar data source and is associated with “alcohol dependence” (see Section 2). Other popular queries were on the subject of: frequency variations, populations and representation of statistical data (biological) obtained for the study.
(3)SELECT allele_freq_id, db_variation_id, pop_db_id, chr_name, position, strand, refallele, refallele_freq, otherallele, otherallele_freq, totalcount FROM snp_allele_freq WHERE position BETWEEN 9200000 AND 9209000
To reduce the amount of data on this query, we decided to fix a search range. Here we show six variations (db_variation_id) included on the position of the chromosome -9.200,000 and 9.209,000-, which indicates that it belongs to chromosome 1 (positive strand) for the Asian population (ASW). In the specific case of the "1009940" variation, this presents allele "G" as reference, which has a frequency of 0.711, and the other allele considered "A", which has a frequency of 0.289 in this population (with a total count of 114 cases) and so on for each variation.
The statistical data, for example, the study of the different calculations related to the LD data -biological-, includes variations and populations. In our next query (4) we give a selection of the most frequently repeated data in the LD table, sorted by the number of repetitions (column "num"):
(4)SELECT pop_db_id, rs_marker1, rs_marker2, Dprime, Rsquare, LOD, fbin, COUNT(*) AS num FROM ld GROUP BY ‘rs_marker1’ ORDER BY num DESC LIMIT 0 , 15
As mentioned in Section 5, "Linkage disequilibrium" (LD) allows us to identify when the genotypes at the two loci are not independent of each other, and relies on statistical models used in population genetics, like: DPrime, Rsquare and LOD (described in Section 5).
(5)SELECT linkaged_id, pop_db_id, rs_marker1, rs_marker2, Dprime, Rsquare, LOD, fbin FROM ld WHERE rs_marker1 = 16919558
This query states, for example, that in "CHD" individuals (Beijing, China) presenting variations "16919558" and "2804311" there is a: DPrime = 0.078, Rsquare and LOD = 0. In this case we can say that these variations applied to the statistical model are not highly dependent on one another, which indicates a probability that there are other variations with a higher complicity.
Thus, by launching query (5) we could see the calculation of LD for the variation (rs_marker1) "16919558" and how this compares with a total of 251 variations (rs_marker2) for the "CHD" population.
In the treatment of this data we developed each step with different control parameters in order to generate a reliable and solid outcome. It is noteworthy that despite the great heterogeneity and dispersion of the data among the different repositories analyzed (genomic data), we can reduce these problems (raw data) by collecting and applying conceptual modeling and data management techniques and thus generate and manage repositories with curated data.
In our research work we seek to exploit this data (haplotypes) in a new way, taking advantage of the current knowledge on genetic variations and themes related to "population genetics”, which positively collaborates in the detection of genetic diseases with personalized attention (personalized medicine).
8 Database evolution according to the conceptual model
In this section we focus on how the way in which the different representations of genomic knowledge are presented affects the structure of the databases used to manage all the data. This is an important part of this work as data is available in different omics data sources and the selected database structure determines how the data is to be managed. The role in this context is to keep an overall conceptual perspective of this data management problem, regardless of the database structure, to obtain a precise conceptual workbench to integrate data correctly.
Figures 9 and 10 show a fragment of a relational database diagram generated from the existing version (previous) and our extended version of the CSHG representing the concepts used in this view.
Our previous version only represents the data loaded into our genomic repository, or variations with a "description". A relational diagram was generated with the "precise" and "imprecise" definitions (with their respective references), omitting the part related to frequencies and populations (not stored).
As mentioned in Sections 2 and 4, haplotypes should be integrated into the conceptual schema because they greatly improve the expressiveness and detail of genetic diagnoses.
Figure 10 shows our extended conceptual schema with the integration of haplotypes. In this case we include all the elements of the variation by "description" and "frequency", generating a more complete relational diagram. This new representation seeks to reduce dispersion issues and prevent redundancy.
This representation means that variations can be treated more specifically, covering all the relevant factors in current genomic knowledge. When the diagram identifies the relationships between variations as part of a haplotype in the genome, the information is enhanced (with a higher level of detail) and points to more accurate results. The fact of incorporating population values into variations plays a key role because of its great impact on the results for end-users (diagnostic / personalized medicine).
This way of representing the data on genome variations defines a new conceptual and logical schema that is very helpful for managing information (loading and handling) and provides a substantial improvement in performance.
Our aim in this section is to show that an accurate representation of the genome data requires precise knowledge of the selected conceptual schema intended to guide the potential different possibilities in which the relevant data can be stored and managed. Although different representations can be made of the same data sources, only a well-defined conceptual schema can provide the unified conceptual perspective needed for managing data correctly and understanding the way in which the data is stored.
9 Lessons Learned and Future Work
The application of conceptual models (CM) to the bioinformatics field is essential for developing Genomic Information Systems (GeIS). Because of recent advances in genomic data repositories, the extension of the previously proposed CSHG is a must. We believe that our conceptual alignment is correct because it extends genetic knowledge in the CSHG through the integration of new concepts related with haplotypes, thus achieving a valid conceptual representation for the different data sources analyzed.
The main contributions of this work are focused on two aspects:
(1) The extension of the conceptual schema (CS), which has great advantages over other representations for its ease of use, understanding and adaptation of genomic data, which are in constant evolution. With our schema we address various gaps identified in the current management of genomic data.
(2) The generation of a new logical schema that adds value in order to analyze, use and exploit the knowledge of haplotypes generated over the years.
One lesson that has been learned is that the application of conceptual modeling [60] techniques contributes to the specification of a sound and reliable schema, to address the various problems detected in genomic data management (see Section 2). The incorporation of conceptual models (CM) into the genomic domain has facilitated data processing, resulting in more powerful "Information Systems" that seek to solve cases related to the large set of heterogeneous data sources (responsible for the high dispersion of data) and incompatible data structures (schemas), among others (see Section 4).
As can be seen in Table 3, when assessing the status of the data associated with haplotypes and frequencies (allelic and genotypic), we find that these repositories provide the information in a wide range of formats, as for example: text (txt) and comma-separated value (csv) files. Firstly, we decided to trace all existing knowledge on this topic; after finishing our conceptual alignment (see Section 6), as an initial step we generated a physical schema to upload and retrieve all the information more efficiently. For this we used the support of database tools such as HeidiSQL [50] and MySQL Workbench [51]. All the data was joined with these tools and then an initial schema was generated (Figure 6) that would display a novel perspective of the data. Figure 6 shows a database schema generated after extracting the different repositories (HapMap, ALFRED, etc.), in which each table gives the source of the data (with the database symbol, e.g. [H]: HapMap).
Finally, after completing this second step of the research methodology, we found that the genomic domain requires the use of approaches that contribute to the organization and management of the "current genomic chaos". With the support of conceptual modeling techniques, the problem is easier to understand and can be solved systematically.
In this work, we developed a conceptual model for the treatment of haplotypes and validated it by determining whether current genomic knowledge (in the different repositories) could be supported by our model. After completing the validation process, we introduced a database into the proposed conceptual model and explained the evolution of the database according to the evolution of the conceptual model.
An additional contribution of this work is the novel conceptual model that combines the knowledge of haplotypes with statistical factors in order to avoid the problems of heterogeneity and the dispersion of data.
Future research work will be aimed at the development of Steps 3 and 4 of the Engineering Cycle (as mentioned in Section 3). The main goal is to check whether current sources could be managed correctly and effectively. Additionally, we will validate the treatment of haplotypes and statistical factors by including this repository in the development of a software tool for genetic diagnosis. Finally, we want to extend the model with studies on the treatment of haplogroups, which include subjects with a similar genetic profile who share a common ancestor.