Abstract
Abstract in Spanish
El melanoma es un tipo de cáncer de piel y es causada por el crecimiento descontrolado de melanocitos atípicos. En las últimas décadas, el diagnóstico asistido por computadora se utiliza para apoyar a los profesionales médicos; Sin embargo, aún no existe una herramienta aceptada a nivel mundial. En este contexto, similar al estado de la técnica se propone un sistema que recibe una imagen dermatoscopica y proporciona un diagnóstico si la lesión es benigna o maligna. Esta herramienta se compone de los siguientes módulos: preprocesamiento, segmentación, extracción de características y clasificación. El preprocesamiento consiste en la eliminación de los pelos. La segmentación es para aislar la lesión. La extracción de características considera la regla ABCD de la dermatoscopía. La clasificación se lleva a cabo por medio de la máquina de vectores de soporte. Los experimentos indican que la propuesta tiene 90.63
Keywords: Melanoma, Automatic Diagnosis, Image Processing, Machine Learning.
Keywords in Spanish: Melanoma, Diagnóstico Automatizado, Procesamiento de Imágenes, Aprendizaje de Máquinas
Received: 2015-11-12 Revised 2016-05-23 Accepted 2016-06-20
DOI: http://dx.doi.org/10.19153/cleiej.19.2.5
The presence of melanocytic skin lesions is becoming increasingly more common, and early detection becomes vital for decision making regarding the treatment to be followed. There are two types of melanocytic lesions: the benign or nevus and the malignant or melanoma. The latter is a skin cancer produced by the uncontrolled growth of melanocytes [1]. This disease could be asymptomatic for the patient, especially in the early stages, so the dermatological inspection is vital [2]. The spread of melanocytes around the body and its asymptomatic nature [3] make it the most lethal skin cancer. In the past years, new methods of non-invasive diagnosis have been developed in order to detect early melanocytic lesions with malignancy features, to reduce unnecessary biopsies, and credit decisions regarding the conduct of the treatment that the specialist suggests [4]. Early diagnosis of melanoma is crucial due to the absence of effective treatments in advanced disease [5].
Melanoma makes up less than 5% of skin cancer cases, but it is the cause of 75% of the deaths that occur by this type of cancer [6]. There is a dramatic increase of melanoma cases; it is estimated that 132,000 cases happen in the world every year, and approximately 66,000 people die because of this, and other types of skin cancer [7]. The determining factors are given by global warming, the continues unprotected exposure to sun rays, and solar beds increases the chances of contracting melanoma [8].
To develop a support tool for diagnosis of non-invasive lesions at early stages with sensitivity and specificity of 89% and 79% respectively [5], using the ABCD rule. The tool, based on techniques of digital image processing and learning machines, should indicate if the image corresponds to a benign or malignant lesion. ]]>
This work is organized as follows: Section 2 presents the concepts of melanocytic lesions and the methods of traditional diagnosis. Section 3 details the tools and algorithms used in the proposed methodology. Section 4 presents the experiments and results obtained. Section 5 shows a discussion of the obtained results. Section 6 shows the final conclusions after the experiments and analysis of the results of the project; and finally, the future works.
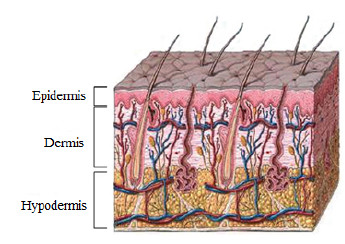
The skin is the largest human-body organ and is composed of three main layers, the epidermis (outer layer of skin), the dermis (inner layer of skin), and the hypodermis (composed of adipocytes that produce and store fat) [1]. In the epidermis there are numerous cells, such as squamous cells, basal cells or cells called melanocytes, as shown in Figure 1. The latter are in charge of producing a substance called melanin, which is the pigment that gives color to the skin [1].
When there is a pigmented skin lesion, there are 2 groups: non-melanocytic lesions, and melanocytic lesions [5]. Melanocytic lesions are classified between benign and malignant lesions, which according to its characteristics may become a melanoma [4].This work focuses on the study of melanocytic lesions due to the mortality characteristic of melanoma.
Melanoma is produced by the uncontrolled growth of cells that accumulate pigment (melanocytes), thus, it can be widely spread on the body through the lymphatic vessels and the blood vessels, giving it its main characteristic of mortality [2].
The factors that cause skin cancer are diverse, however, 86% of melanomas is due to the exposure to ultraviolet radiation (UV) from the Sun [12]. There are a number of factors that increase the risk of developing melanoma [13] such as: the presence of atypical nevus, the presence of nevus exceeding 40 units, the presence of congenital nevus, having a family history of atypical nevus, of melanoma, or to have been exposed for many hours to ultraviolet rays (Sun, UVB, UVA) even though having a completely tanned skin.
The dermoscopy is a non-invasive technique of imaging diagnosis which is used for the observation of skin lesions, allowing the visualization of the structures of the epidermis and dermis [14]. An instrument called a dermatoscope is used for this technique, which consists of a light source and an image amplification system [8]. Figure 2 (a) shows an image taken with a regular camera, while Figure 2 (b) shows an image taken with the dermatoscope, offering a better view of the characteristics of the area of interest.
The use of this technique reaches 89% of sensitivity, and 79% of specificity of the professional [16, 17]. This is the reason why dermoscopy in the diagnosis of the lesions is of vital importance in little expressive lesions and at early stages [16, 17].
The diagnostic method commonly accepted in Dermatology is the 2-stage analysis, where the first stage analyses the lesion of the skin to determine whether it is melanocytic or non-melanocytic; and then the lesion is analyzed to see if the melanocytic one is malignant or benign (second stage) [18, 3, 19].
In the first stage, it is important to detect noticeable features of the melanocytic lesions (for a better study, consult in [14] and [20]). In the second stage, exhaustive and thorough methods of analysis [5] are required for the diagnosis of benign or malignant lesion. There are several methods commonly used, such as: the Pattern Analysis in Zalaudek et al. [21], the Menzies Method in Menzies et al. [22], the 7-point checklist in Argenziano et al. [23], and the ABCD rule in Stolz et al. [24], the latter is used in this work.
Developed by Stolz [24] in 1993, it analyzes 4 criteria: asymmetry, borders, colors, and dermoscopic structures. Asymmetry is the pattern generated by the uncontrolled growth of the lesion, it is measured by dividing the pigmented lesion into 2(two) axes, looking for the best symmetry possible [5, 25]. Figure 3 shows a symmetric and an asymmetric lesion with 2 (two) marked axes. The first axis is outlined on the longest part of the lesion and is called the main axis, the second axis is outlined perpendicular to the main axis.
Borders are patterns associated with abnormal terminations of color that melanocytic lesions have, i.e., borders have color variations. The area of interest is divided into 8 segments. Figure 4 shows normal borders indicated with an X, the others are considered as abrupt.
Dermoscopic structures taken into account are detailed below:
Some state-of-the-art reported works are presented next, that propose automated diagnostic systems. ]]>
In Batugo [1] an automated diagnostic system was proposed for the detection of melanomas, composed by the following modules: The pre-processing, which consists of the elimination of borders or margins that should not be present in the image at the time of the analysis. Also seeks to isolate objects foreign to the lesion, such as hair. The proposed segmentation was performed by means of the Clustering Method using Gaussian mixtures; the feature extraction was performed based on the ABCD rule. Classification was carried out using linear discriminant classifier [26], in order to obtain the diagnosis using the previously extracted characteristics. The tests were carried out with 100 images whose diagnosis is detailed as follows: 29 melanomas and 71 nevus. The evaluation of the system obtained a 79.31% of sensitivity, and 71.83% of specificity.Similar to the previous work, Oliveira [27] developed a diagnostic system of pigmented lesions formed by 4 modules: pre-processing of images was carried out by anisotropic diffusion filter [28] to eliminate the noise. The segmentation module was carried out by using the Chan Vese Method [29]. The feature extraction was performed using the ABC-T (Assimetry, Border, Color, Texture) rule. The last characteristic was added in order to consider the ”lesiones queranocíticas” (Keratinocytic Lesions) which are anomalies of the skin that appear as a crust on its surface. Classification was carried out using the SVM (Support Vector Machine) classifier [29]. The evaluation according to the sensitivity metrics and specificity were of 73.81% and 76.67% respectively. The image bank was composed of 408 digital images, of which 62 were nevus, 86 images with keratosis and 260 melanomas.
Em Rahman et al. [30] combined different classifiers in order to develop a melanoma automatic recognition system using dermoscopic images. For the segmentation, the clustering algorithm was used, Fuzzy C-means [31].Then, an opening and a morphological closing were applied to eliminate noise and soften the border, obtaining this way the outline of the lesion. For the classification of the skin lesions, the combination of the technology of the SVM, GML [32], and K-NN [33] classifiers were used. The following results were obtained: 62.50% of specificity and 83.75% of sensitivity, from a total of 358 dermoscopic images using 40% for training and 60% for tests.
In Alcon et al. [34] a diagnostic system of pigmented lesions was proposed. Segmentation was carried out using the Otsu method [35]. The characteristics were extracted through the ABCD method, totaling 55 characteristics. The classification was carried out using the K-NN classifier. Tests conducted on the total of 152 digital images, yielded the following results: 45 nevus and 107 melanoma, corresponding to a 68% of specificity, and 94% of sensitivity.
It is presented in Celebi et al. [36] a methodological approach for the classification of pigmented lesions of the skin using dermoscopic images. Median filter is used for pre-processing and Otsu’s method for segmentation. In the feature extraction, the following characteristics were used: the area of the lesion, relation of appearance, asymmetry, compactness. For the classification, they used SVM and in experiments conducted on a set of 564 images, they produced a specificity of 92.34% and a sensitivity of 93.33%.
It is important to note that there are several related works that do not follow any dermatological rule to find the diagnosis, since they propose the study of characteristics such as color and texture of the lesion in a specific way, as the ones reported in [37, 38, 39, 40, 41, 42]. In Scharcanski et al. [43] you can see a summary of the work using computer vision techniques in the diagnosis of skin cancer and provides an orientation of medical image analysis.
This topic is controversial in terms of whether to use or not to use the diagnostic methods approved by the dermatologists [37]. In this context, this work prefers to follow the line accepted by them, as it is the ABCD rule, developing techniques which comply with the requirements of the dermatologic community and thus improve the performance reported in the state-of-the-art.
Based on the work of the state-of-the-art, 4 main modules are proposed: Pre-processing, Segmentation, Feature Extraction, and Classification.
An 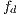 image (dermoscopic melanocytic image) for the removal of hairs is received, to which the bottom-hat operator is applied on each channel of the RGB space. Then, the maximum value among the 3 channels is taken, as shown in Figure 7 (b). Then, the Otsu algorithm [35] is applied to make the image binary, the result of this operation can be seen in Figure 8 (a), and it is used as a mask for painting the hairs with a very different color from that of the lesion, in this case, green, as shown in Figure 8 (b). Finally, Inpainting is applied, proposed by [44], which is an algorithm of image restoration that uses information from the rest of it to rebuild the areas associated with the hairs. The output of this module is an
image (dermoscopic melanocytic image) for the removal of hairs is received, to which the bottom-hat operator is applied on each channel of the RGB space. Then, the maximum value among the 3 channels is taken, as shown in Figure 7 (b). Then, the Otsu algorithm [35] is applied to make the image binary, the result of this operation can be seen in Figure 8 (a), and it is used as a mask for painting the hairs with a very different color from that of the lesion, in this case, green, as shown in Figure 8 (b). Finally, Inpainting is applied, proposed by [44], which is an algorithm of image restoration that uses information from the rest of it to rebuild the areas associated with the hairs. The output of this module is an 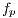 image, as shown in Figure 9, which at the same time is the entrance to the next module (segmentation).
image, as shown in Figure 9, which at the same time is the entrance to the next module (segmentation).
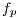 image in grayscale Figure 10 (a). To remove the peaks of intensity between pixels, median filter is applied with a window size of 4 x 4. The level of grey of each point is replaced by the value of gray of a point by the average of the levels of grey of a certain vicinity, the resulting image is shown in Figure 10 (b). The adjustment of the intensity values of the image is made using the value of gamma = 0.01, in order to reduce noise, as shown in Figure 11 (a).
image in grayscale Figure 10 (a). To remove the peaks of intensity between pixels, median filter is applied with a window size of 4 x 4. The level of grey of each point is replaced by the value of gray of a point by the average of the levels of grey of a certain vicinity, the resulting image is shown in Figure 10 (b). The adjustment of the intensity values of the image is made using the value of gamma = 0.01, in order to reduce noise, as shown in Figure 11 (a).
To improve the contrast of the image so that the lesion can be highlighted, the algorithm of improvement of contrast (Contrast Limited Adaptive Histogram Equalization) CLAHE is used [45], with Clip Limit = 0.01 and with contextual regions equal to 2 x 2 (see Figure 11 (b)). Once the image is improved, the threshold is calculated with the maximum-entropy automatic thresholding algorithm [45] and the binary mask resulting from this stage can be seen in Figure 12 (a). The image of the lesion associated to the binary mask can be seen in Figure 12 (b).
This work uses the ABCD rule for the diagnosis of Melanoma. In that sense, the features to be extracted are: Asymmetry, Border, Color, Reticular Pattern, Dots and Glabules, Structureless Areas, and Pseudopods.
This feature is obtained through the asymmetry index which is calculated using a combination of instructions. This index indicates asymmetric is the lesion, i.e., the greater the value of it, the more asymmetric the lesion is; and the lower the value of it, the more symmetrical the lesion is. The segmented lesion shown in Figure 13 (a), is rotated through its centroid by bi-linear interpolation method [24], and is shown in Figure 13 (b).

Once the lesion is rotated it is then divided through the main axis, as shown in Figure 14 (a) and then divided by the secondary axis, in Figure 14 (b) there is the lesion divided by the secondary axis.
The asymmetry index 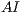 is calculated by the expression (1)
is calculated by the expression (1)
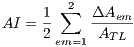 | (1) |
where 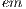 = 1 is the major axis and
= 1 is the major axis and 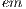 = 2 is the minor axis, is the area of the non-overlapping lesion; and
= 2 is the minor axis, is the area of the non-overlapping lesion; and 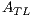 is the area of the lesion. In this way, the first feature is obtained and is denoted by
is the area of the lesion. In this way, the first feature is obtained and is denoted by 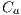 .
.
For the border analysis, the lesion is divided into 8 regions. Each retrieved sector, as shown in Figure 15, is analyzed separately. In this way, if the color of the lesion varies from the center towards the border, it is said that it has an abrupt border, otherwise is a normal or regular border, lesions with abrupt termination are associated with melanoma. The color variation in the borders is measured by calculating the variance from the center of the lesion until the border of it, where high values of variance indicates a border with abrupt termination, and low values indicate a normal border.
In this way, the feature of border is obtained and is denoted by 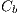 .
.
In the study of this feature, 6 colors are taken into account: light brown, dark brown, blue-gray, red, black and white.The presence of these colors suggest melanoma. Each color is detected by means of the Euclidean distance of each pixel to the corresponding intensities to that color. Table 1 shows the value of intensity of each color. The lower the value of the calculated distance suggests closeness to a color. Figure 16 (a) shows a lesion with the presence of the blue-gray color, and in Figure 16 (b), a lesion that contains the colors dark brown and light brown. ]]>
The presence of different colors in the lesion points to a positive diagnosis of Melanoma. For the presence of the color, the obtained features are: light brown denoted by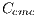 , dark brown denoted by
, dark brown denoted by 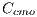 , blue-gray denoted by
, blue-gray denoted by 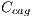 , red denoted by
, red denoted by 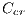 , black denoted by
, black denoted by 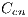 and white denoted by
and white denoted by 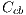 , represented by the vector
, represented by the vector 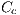 , which in turn is given by:
, which in turn is given by: ![Cc = [Ccmc,Ccmo,Cag,Ccr,Ccn,Ccb]c](/img/revistas/cleiej/v19n2/2a0617x.png)
They are different patterns associated with melanoma, it includes the study of 5 structures: Linear Branches denoted by 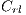 , Atypical Pigmented Reticulum denoted by
, Atypical Pigmented Reticulum denoted by 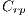 , Structureless Areas denoted by Cad, points denoted by
, Structureless Areas denoted by Cad, points denoted by 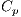 , and globules
, and globules 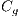 . This set of features is represented by the
. This set of features is represented by the 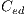 vector defined by:
vector defined by: ![Ced = [Crl,Crp,Cad,Cp,Cg]ed](/img/revistas/cleiej/v19n2/2a0624x.png) . Details of the recognition process of these features are given below.
. Details of the recognition process of these features are given below.
Linear Branches: Pseudopods are bulbous and bent projections as shown in Figure 18 (a), the borders are finger-like structures (in the form of fingers).
To detect the pseudopods, the variances of the Euclidean distance are analyzed between the centroid and the border of the lesion.The binary mask from the original image can be seen in Figure 18 (b). To this mask, the morphological operator dilation is applied with a structuring element with the shape of a disc of radious 1, Figure 18 (c). The border of the lesion results from the subtraction of the dilated image and the original image, as shown in Figure 18 (d). The variance is calculated as a measure of dispersion of the distances to the border, a high value suggests the presence of pseudopods.
Atypical Pigmented Reticulum:
A relevant feature associated to the diagnosis of melanoma is the atypical pigmented reticulum. Figure 19 (a) shows a lesion with cross-linked, to diagnose as typical or atypical, a series of steps will be taken, which are explained below.
On the image of the segmented lesion in grayscale, Figure 19 (b), it is applied the detection of border with an Laplacian-of-Gaussian operator (LOG) [46], the result is shown in Figure 19 (c). The cross lines which form a mesh over the lesion; and, also the outside border of the lesion that must be removed, can be observed. With the aim of obtaining only formed holes, the border is calculated and then it is subtracted from Figure 19 (c).
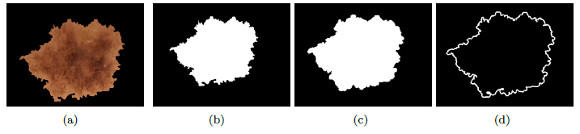


Structureless Areas: Structureless areas are regions of the lesion to which the aforementioned structures can not be remunerated.
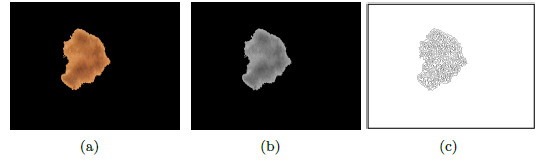
In the center of the Figure 21 (a) a lesion is shown, with a structureless area of a dark brown color, indicated by the red square. The image of the lesion is in grayscale as shown in Figure 21 (b). This lesion is segmented by the traditional Otsu algorithm, to isolate the dark areas as can be seen in Figure 20 (c). Then, the region that belongs to the structureless area gets separated, and the resulting image is shown in Figure 21 (d).
Dots and Globules: Dots and globules are differentiated by their size, the first are almost invisible to the naked eye. Figure 22 (a) shows a lesion presenting various dark-brown globules. ]]>
With the aim of locating globules, first, an improvement of the distribution of the pixels of the image in grayscale is made (Figure 22 (b)), using the average filter with a window size of 4 x 4, as shown in Figure 22 (c). To highlight them, an intensity adjustment is done with gamma = 0.01 and a contrast improvement is applied through the equalization of the histogram using the CLAHE method with Clip Limit = 0.01 and contextual regions equal to 2 x 2, as shown in Figure 22 (d).

The binary image is obtained through the Otsu method (Figure 23 (a)). Then, closure is applied with a structural element is the shape of a disk and size 3, in such a way that the nearby components are united as can be seen in Figure 23 (b). The vector of 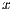 characteristics consisting of:
characteristics consisting of: ![x = [Ca;Cb;Cc;Ced ]](/img/revistas/cleiej/v19n2/2a0626x.png) where each element represents the sub-vectors previously calculated.
where each element represents the sub-vectors previously calculated.
In this work, the classification of lesions will be done by using the SVM classifier (Support Vector Machine) [29]. Entry data will be formed by 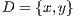 where
where 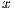 is the set of vectors of features while
is the set of vectors of features while 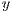 is the set of labels. They represent one of the two classes,
is the set of labels. They represent one of the two classes, 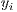 =0 for benign lesions and
=0 for benign lesions and 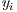 =1 for malignant lesions associated with the i-th melanocytic lesion.
=1 for malignant lesions associated with the i-th melanocytic lesion.
The classification will have two stages: a) The construction of the classifier for the learning of the parameters of the system, this process is performed with a set of training. b) The system’s tests to evaluate the classifier’s performance is carried out with a set of tests that are independent from the set of training.
This section presents the evaluation metrics used, the experiments done, the results obtained, and those of the state-of-the-art; then, there is a discussion about the results.
120 dermatoscopy images were used for the tests (non-melanocytic and melanocytic) acquired through a dermatoscope computerized video with polarized light, which uses lenses with 20 to 70x given by the Centre for Processing Digital Images of the Faculty of Sciences, Universidad Central de Venezuela [47].
The images were analyzed by Dr. Lizza Salgueiro, from the Department of Dermatology of the Universidad Nacional de Asunción, since they were not previously diagnosed. According to her diagnosis, 16 non-melanocytic and 104 melanocytic lesions were found, from which 76 were malignant lesions, and 28 were benign lesions.
The tests that were carried out for the module of hair removal, consists of the application of 2 techniques for its comparison.
Figure 24 (a) shows a lesion, with the presence of hair, both, on the lesion as well as on the skin; the goal is to perform the hair removal using the following methods:
The first approach is based on the following operations [47] : ]]>
First, it applies morphological filters. Then, the Botton-Hat operator is used. Through segmentation by thresholding, the hairs from the skin are separated. Finally, the associated areas with the hairs are reconstructed with the average filter.The second technique, proposed in this work (section 3.1.1), is based on: Obtaining a mask of hairs through the Botton-Hat operator and the traditional Otsu algorithm. On this mask, the Inpainting algorithm [44] is applied for the replacement of the pixels associated with hairs.
Figure 24 (b) shows the resulting image from the first method previously mentioned, it can be noticed that the elimination of hair affected some areas of the skin and the lesion, which is not recommendable. Finally, Figure 24 (c) shows the image without hairs, resulting from applying the second method; it can be noticed that the colors of the lesion remained intact.
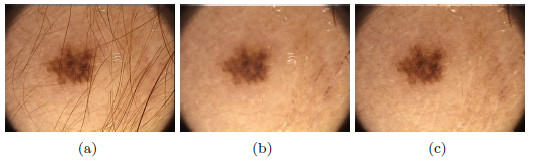
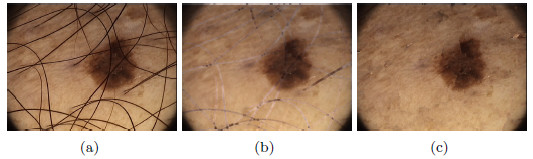
As a second test, Figure 25(a) shows a lesion with presence of hairs with greater thickness than the previous example. Figure 25(b) shows the image resulting from the first method, and it can be noted that eliminating the hairs affected the area of the lesion, and some whitish markings with the shape of hairs were left. Figure 25(c) shows the hairless image resulting from applying the Inpainting algorithm, the lesion with a total absence of hairs, unmarked and without alteration of the colors, can be noticed in the image. These methods were visually evaluated by the professional dermatologist. After several tests performed on lesions with thick and fine hairs, the expert arrived to the conclusion that the thickest hairs are best removed with the second method exposed. Despite the computational cost of the Inpainting algorithm, as it is mentioned in [44], in this work is used due to the improvement it represents visually in the results. ]]>
Segmentation seeks to isolate the lesion from the rest of the skin for further study. Performance metrics were defined in this section to evaluate the segmentation method. Four possible outcomes can be obtained from a segmented lesion:
In Figure 26 you can see the representation of TP, FN, TN and FP variables regarding the segmented lesion and its ideal segmentation, where the pixel associated with the targeted zone is given by 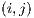 , the ideal image represents the segmentation done by a professional, and the segmented image represents the segmentation of the lesion obtained automatically.
, the ideal image represents the segmentation done by a professional, and the segmented image represents the segmentation of the lesion obtained automatically.
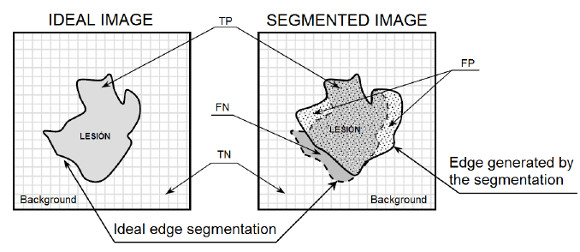
The metrics used are:
Currently, there are several segmentation methods for melanocytic lesions due to the important role that they play in a diagnostic system. In [49] a comparison of 6 segmentation methods applied to melanoma were carried out, where it was concluded that the AS (Adaptive Snake) and EM-LS (Expectation Maximization Level Set) methods were strong and useful for the segmentation of the lesion in a computer-aided diagnostic system in order to help the clinical diagnosis of dermatologists. On the other hand, this work has as its main objective the improvement of the performance of final diagnosis for which 2 automatic segmentation algorithms were tested: the Otsu’s traditional algorithm, and that of Maximum Entropy. Table 2 shows the results of the 2 algorithms. The results obtained were compared with images of ideal segmentation made by the dermatologist.
As shown in Table 2, the results are similar in terms of performance, specifically, for pigmented lesions were reported [8] which is convenient to use the method of maximum entropy, so that in this work such algorithm is used for the segmentation.
The objective of the classifier on the proposed methodology is to return a negative or positive diagnosis, ie, whether the input samples represent a malignant or benign lesion. For a given sample, a diagnostic system can lead to one of four possible outcomes:
In Table 3 the matrix that includes the four possible outcomes is shown. The Sensitivity, Specificity and Accuracy metrics are taken into account in order to assess the results.
Table 4 shows the formal definition of the performance metrics for specificity, sensitivity and accuracy.
For the construction of the classifier and the making of the experimental testing, the Bioinformatics Toolbox for Matlab R2013b was used. Experiments are explained below.
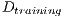 and the second is called test set
and the second is called test set  . Then, the set of training is formed, which contains 32 images of which were randomly selected 12 benign and 20 malignant with a probability p = 30 without repetition.The rest of the images (72 images) are used for the validation of the classifier obtained in the previous step. The results of this process are shown in Table 5.
. Then, the set of training is formed, which contains 32 images of which were randomly selected 12 benign and 20 malignant with a probability p = 30 without repetition.The rest of the images (72 images) are used for the validation of the classifier obtained in the previous step. The results of this process are shown in Table 5. It is necessary to use other methods since in experiment I the minimum standard of the Specificity Metrics was not reached (greater than 79%). The cross-validation is used to give strength to the classifier. For the K-Fold Cross-Validation, the data set is divided into K groups and K iterations are performed. Each iteration uses a group for the test 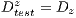 and the K-1 remaining groups
and the K-1 remaining groups 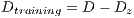 are used for training with
are used for training with 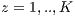 . In pseudocode 1, the collection of the
. In pseudocode 1, the collection of the 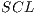 classification system is described, whereas in the results gathered from this process are shown in Table 6.
classification system is described, whereas in the results gathered from this process are shown in Table 6.
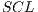 Classification System
Classification System This data partition is unworkable when there is small amount of data, since the amount of images of training that are used by each group still cannot provide a classifier that complies with the performance standards of the dermatoscopy that are sought in this work. Note that the performance by group is less than that obtained by the retention method.
Due to the unfavorable results in experiment II (shown in Table 6 - less than the standards of specificity and sensitivity of 79% and 89%, respectively) Random Cross-Validation was conducted where K iterations were made, but by each iteration, a set of test 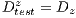 and the set of training
and the set of training 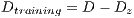 . must be chosen randomly.
. must be chosen randomly.
In the k-th iteration, it is calculated the k-th classification system CL to conform classifiers K, progressively. In this context, it is in the K-th iteration, the K-th classification system 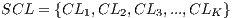 whose performance is obtained from the average of the classifiers K. Next, The Algorithm 2 is shown, which describes the collection of the classification system
whose performance is obtained from the average of the classifiers K. Next, The Algorithm 2 is shown, which describes the collection of the classification system 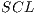 based on the Random Cross-Validation.
based on the Random Cross-Validation.
As shown in Figure 27, the results begin to stabilize from iteration K = 12, so 3 iterations more are performed, until reaching a K=15.
The final average of such tests can be seen in Table 7 that corresponds to a 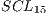 system.
system.
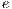 iterations are performed, where
iterations are performed, where 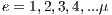 and each assembled system
and each assembled system 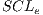 , obtained through routine SSVM, it is evaluated by the metric of accuracy.The assembled system selected will be the one that has obtained greater accuracy.
, obtained through routine SSVM, it is evaluated by the metric of accuracy.The assembled system selected will be the one that has obtained greater accuracy. 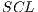
For the specific problem 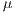 = 30 was used, from the 30 iterations, the assembled system used in the experiments III was selected.
= 30 was used, from the 30 iterations, the assembled system used in the experiments III was selected.
In operation time, an assembled system consisting of classifiers K, obtains the final diagnostic by means of simple vote, because the output of each classifier CL is a discrete value.
The results of the assembled system of the experiment III in operational time are shown in Table 8.
Notice the good results obtained, but they can be tricky since all the data is used to train the classifiers. For this reason, in step A, a set of the total data is isolated for the validation of the assembled system , and step B takes the rest of the set for training and testing, having as base the method of random cross-validation. This experiment is performed with two different data sets, the first set is using the ideal segmentation made by the professional, and the second using the proposed segmentation; both results are shown in Table 9.
This experiment is carried out in order to observe the impact of a non-ideal segmentation on the system, which consists in building a classifier based on images with ideal segmentation (generated by the medical professional). All the images are used for the test data, but they follow the proposed segmentation process of this work.The results obtain are shown in Table 10.
This section presents the discussions on the results obtained from the experimental tests of the segmentation and classification modules.
It must be taken into account that segmentation can be affected by the method of hair extraction used in each work, which could lead to unfavorable or advantageous results.
The results obtained in the segmentation made on the same bank of images, by means of the proposed method obtained 97% of accuracy as shown in Table 11; in the same way, it can be observed that the results of [47] were favorable but below the proposal. This could be due to the hair extraction method used in this work, hence the importance of a good method for hair extraction.
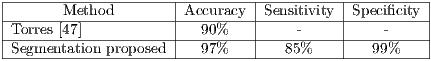 47]-----------|---90%---|----------|----------| -Segmentation-proposed-----97%-------85%--------99%----- " >
47]-----------|---90%---|----------|----------| -Segmentation-proposed-----97%-------85%--------99%----- " >Prior to the discussion of the results, it is important to clarify that the bases of images used in this work and those of the state-of-the-art are different because there are no public database due to confidentiality reasons kept between doctors and their patients. Regarding the reproduction of such works, they could not be done because of the scarce information provided by the authors, in most of the cases, the main obstacle was the lack of information on the methods used for the extraction of hairs. It could also be observed that the characteristics used in these works require knowledge of the consecutive acquisition protocol for image taking, information that is not available, and it is the reason why features such as the area and the diameter of the lesion could not be used. For these reasons, the proposed scheme is incomparable with the state-of-the-art approaches, it can only be alleged that the results obtained in this work are well positioned as shown in the Table 12.
]]>
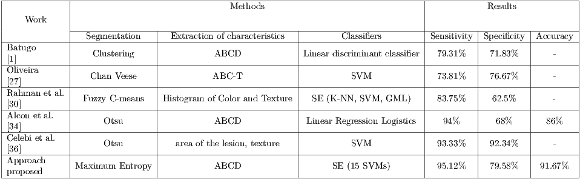 1]-----------|-----------------|----------------------------|-------------------------|----------|----------|---------| | Oliveira | Chan Veese | ABC -T | SVM | 73.81% | 76.67% | - | |-[27]----------|-----------------|----------------------------|-------------------------|----------|----------|---------| | Rahman et al.| Fuzzy C-means |Histogram of Color and Texture SE (K -NN, SVM, GML ) | 83.75% | 62.5% | - | |-[30]----------|-----------------|----------------------------|-------------------------|----------|----------|---------| | Alcon et al. | | | | | | | |-[34]----------|------Otsu-------|----------ABCD--------------|-Linear Regression-Logistics-|---94%----|---68%----|--86%----| | Celebi et al. | | | | | | | |-[36]----------|------Otsu-------|---area-of the lesion, texture-|----------SVM------------|--93.33%---|-92.34%---|---------| | Approach | | | | | | | --proposed-------Maximum--Entropy------------ABCD---------------------SE-(15 SVMs-)---------95.12%-----79.58%-----91.67%---" >
1]-----------|-----------------|----------------------------|-------------------------|----------|----------|---------| | Oliveira | Chan Veese | ABC -T | SVM | 73.81% | 76.67% | - | |-[27]----------|-----------------|----------------------------|-------------------------|----------|----------|---------| | Rahman et al.| Fuzzy C-means |Histogram of Color and Texture SE (K -NN, SVM, GML ) | 83.75% | 62.5% | - | |-[30]----------|-----------------|----------------------------|-------------------------|----------|----------|---------| | Alcon et al. | | | | | | | |-[34]----------|------Otsu-------|----------ABCD--------------|-Linear Regression-Logistics-|---94%----|---68%----|--86%----| | Celebi et al. | | | | | | | |-[36]----------|------Otsu-------|---area-of the lesion, texture-|----------SVM------------|--93.33%---|-92.34%---|---------| | Approach | | | | | | | --proposed-------Maximum--Entropy------------ABCD---------------------SE-(15 SVMs-)---------95.12%-----79.58%-----91.67%---" >The Retention Method was used in experiment I, where it can be observed that there is an unbalance of loads between benign and malignant lesions, being the latter the kind which prevails. In these cases, the majority of classifiers focuses on learning from the large kinds resulting in a poor accuracy of classification for the smaller kinds. This is verified in the results obtained in this experiment being specificity lower than sensitivity, it is worth mentioning that the result obtained in sensitivity is crucial because the proper detection of malignant lesions is a priority.
With the aim of improving specificity, experiment II was conducted, whose results were not favorable even affecting sensitivity and showing the effect of insufficient data. This happened because of the division into sets of training and testing of equal size in order to obtain different classifiers which led to few images to train each classifier, which in turn, led to conduct experiment III.
In experiment III encouraging results were obtained, with a slight improvement in specificity and without sacrificing sensitivity, this is the effect of using more benign lesions during training.
With the aim of providing strength to the proposed method, an assembled system was used, which consisted of 15 classifiers.First of all, the assembled system of the experiment was formed but it was concluded that these results suggest over-training, which could lead to a poor performance for new samples.
For this reason, experiment IV was conducted, where the results are satisfactory for an unknown validation group.
Considering the sensitivity and specificity of the dermoscopy as a basis, the proposed scheme is valid since it reached these metrics which constitute the general objective of this work.
It can be observed in experiment V that non-ideal segmentation degrades the performance of the classifier that was built with all of the ideal images, getting a more negative impact on specificity than on sensitivity. With this, we can conclude that work must continue in the improvement of a segmentation algorithm that can reach to be as ideal as it can get.
A set of image processing techniques were used to obtain features of the ABCD rule, quite used in dermatology, and use it as an input to a classifier.This is done in order to be able to give a diagnosis of skin cancer, specifically, melanoma. Several tests were conducted while classifying, so as to evaluate its performance and get acceptable results by dermatology experts. According to the performance metrics, very favourable results were obtained, taking into account the state-of-the-art approaches. However, for a work confrontation the same database should be used, which was not possible due to lack of public databases. In Dermatology, it is considered as a valid tool only if it reaches the performance of the diagnosis using the dermatoscopy, so the scheme proposed constitutes as a valid tool for the dermatologist. The proposed scheme complies with the outlined objectives, the generals and the specific ones. The tool has been developed in order to assist the dermatologist and not to replace them; it is intended to contribute to obtain an objective diagnosis, independent from the professional experience.
As future work, the authors propose to carry out the feature extraction using another rule, either pattern analysis [21], the Menzies method [22] or another. To build a tool that will point out each of the features found in the lesion and that will display itself as a means of information for the medical professional. To implement a method for the extraction of hairs with lower computational cost and implement metrics to measure its performance. To do tests with a standard database, with a defined acquisition protocol in such a way that it can include characteristics like the area and the compactness of the lesion. ]]>
[1] G. A. Batugo, “Reconocimiento automático de melanomas mediante técnicas de visión por ordenador y reconocimiento de patrones,” Universidad Carlos III de Madrid, Tech. Rep., 2013.
[2] A. Sboner, C. Eccher, E. Blanzieri, P. Bauer, M. Cristofodolini, G.Zumiani, and S.Forti, “A multiple classifier system for early melanoma diagnosis,” Artificial Intelligence in Medicine, vol. 27, no. 1, pp. 29–44, 2003. [Online]. Available: http://dx.doi.org/10.1016/S0933-3657(02)00087-8
[3] P. Braun, H. Rabinovitz, M. Oliviero, A. Kopf, and J. Saurat, “Dermoscopy of pigmented skin lesions,” Journal of the American Academy of Dermatology, vol. 48, no. 5, pp. 679–693, may 2003. [Online]. Available: http://dx.doi.org/10.1016/j.jaad.2001.11.001
[4] Parikh and Hitesh, “A survey on computer vision based diagnosis for skin lesion detection,” International Journal of Engineering Science and Innovative Technology, vol. 2, no. 2, pp. 431–437, 2013.
[5] P. Zaballos, C. Carrera, S. Puig, and J.Malvehy, “Criterios dermatoscópicos para el diagnóstico del melanoma,” Medigraphic, vol. 32, 2004.
[6] La Organización Mundial de la Salud desaconseja el uso de camas solares a las personas menores de 18 años. http://www.who.int/mediacentre/news/notes/2005/np07/es/. Accessed: 2016-06-27.
[7] P. Ramos, F. C. nete, R. Dullak, L. Bolla, N. Centurión, A. Centurión, S. Chamorro, A. Chaparro, and F. Chaves, “Epidemiología del cáncer de piel en pacientes atendidos en la cátedra de dermatología de la facultad de ciencias médicas de la universidad nacional de asunción, paraguay (2008-2011),” ANALES de la Facultad de Ciencias Médicas, vol. 45, no. 2, pp. 49–69, 2012.
]]> [8] O. Blandom, “A support tool for melanoma diagnosis by using dermoscopy images,” Master’s thesis, Universidad Nacional de Colombia, Manizales, Colombia, 2010.[9] H. Zhou, G. Schaefer, M. Celebi, H. Iyatomi, K. Norton, and F. L. T. Liu, “Skin lesion segmentation using an improved snake model,” Engineering in Medicine and Biology Society (EMBC), 2010 Annual International Conference of the IEEE, 2010. [Online]. Available: http://dx.doi.org/10.1109/IEMBS.2010.5627556
[10] H. Ganster, A. Pinz, R. Rohrer, E. Wildling, M. Binder, and H. Kittler, “Automated melanoma recognition,” Medical Imaging, IEEE Transactions on, vol. 20, no. 3, pp. 233–239, 2001. [Online]. Available: http://dx.doi.org/10.1109/42.918473
[11] A. Lifshitz. Llega Zelboraf: una medicina nueva contra el melanoma, un cáncer de la piel. http://www.vidaysalud.com/diario/cancer/llega-zelboraf-una-medicina-nueva-contra-el-melanoma-un-cancer-de-la-piel/. Accessed: 2016-06-27.
[12] Melanoma. http://www.dmedicina.com/enfermedades/cancer/melanoma. Accessed: 2016-06-27.
[13] R. J. Friedman, D. S. Rigel, A. W. Kopf, and D. Polsky. Melanoma. http://cancerdepiel.org/cancer-de-piel/melanoma. Accessed: 2016-06-27.
[14] M. Fossati, C. Guebenlián, M. Restano, and A. Wolf, “Lesiones melanocíticas,” Dermatoscopía Elemental, 2013.
[15] H. P. Soyer, R. Hofmann-Wellenhof, R. H. Johr et al., Color atlas of melanocytic lesions of the skin. Springer, 2007.
[16] M. de Troya Martína, N. B. Sáncheza, I. F. Canedoa, M. F. Eliceguia, R. F. Liébanab, and F. R. Ruizc, “Estudio dermoscópico del melanoma maligno cutáneo: análisis descriptivo de 45 casos,” vol. 99, no. 1, pp. 44–53, 2008.
[17] I. Stanganelli. Dermoscopy. http://emedicine.medscape.com/article/1130783-overview. Accessed: 2016-06-27.
]]> [18] G. Argenziano, H. Soyer, S. Chimenti, and G. Ruocco, “Dermoscopy of pigmented skin lesions,” European Journal of Dermatology, vol. 11, no. 3, pp. 270–277, may 2002.[19] Malvehy and Puig, “Principios de dermatoscopía,” 2002.
[20] A. Bazzano, M. Cueto, C. García, and J. Pérez, “Lesiones no melanocíticas,” Dermatoscopía Elemental, 2013.
[21] I. Zalaudek, G. Argenziano, H. Soyer, R. Corona, F. Sera, A. Blum, R. Braun, H. Cabo, G. Ferrara, A. Kopf et al., “Three-point checklist of dermoscopy: an open internet study,” British journal of dermatology, vol. 154, no. 3, pp. 431–437, 2006. [Online]. Available: http://dx.doi.org/10.1111/j.1365-2133.2005.06983.x
[22] S. W. Menzies, C. Ingvar, and W. H. McCarthy, “A sensitivity and specificity analysis of the surface microscopy features of invasive melanoma,” vol. 6, no. 1, pp. 55–62, 1996.
[23] G. Argenziano, G. Fabbrocini, P. Carli, V. D. Giorgi, E. Sammarco, and M. Delfino, “Epiluminescence microscopy for the diagnosis of doubtful melanocytic skin lesions comparison of the abcd rule of dermatoscopy and a new 7-point checklist based on pattern analysis,” JAMA Dermatology, vol. 134, no. 12, pp. 1563–1570, December 1998.
[24] Stolz, W. Riemann, A. Cognetta, A. Pillet, L. Abmayr, and W. Hoelzel, “Abcd rule of dermatoscopy: a new practical method for early recognition of malignant melanoma.” European Journal of Dermatology, vol. 4, no. 7, pp. 521–527, 1994.
[25] R. Montero, A. H. Abúndez, and A. Zamarón, “Descripción de la asimetriá de neoplasias en piel utilizando el concepto de compacidad,” IX Congreso Internacional sobre Innovación y Desarrollo Tecnológico CIINDET 2011, Cuernavaca Morelos, México, 2011.
[26] C. Pérez López, “Métodos estadísticos avanzados con spss,” Thompson. Madrid, 2005.
[27] R. B. Oliveira, “Método de detecção e classificação de lesões de pele em imagens digitais a partir do modelo chan-vese e máquina de vetor de suporte,” Universidade Estadual Paulista Júlio de Mesquita Filho, Tech. Rep., 2012.
]]> [28] C. A. Z. Barcelos, M. Boaventura, and E. C. Silva Jr, “A well-balanced flow equation for noise removal and edge detection,” Image Processing, IEEE Transactions on, vol. 12, no. 7, pp. 751–763, 2003. [Online]. Available: http://dx.doi.org/10.1109/TIP.2003.814242[29] C. Cortes and V. Vapnik, “Support-vector networks,” Machine learning, vol. 20, pp. 273–297, 1995. [Online]. Available: http://dx.doi.org/10.1007/BF00994018
[30] M. Rahman, P. Bhattacharya, and B. Desai, “A multiple expert-based melanoma recognition system for dermoscopic images of pigmented skin lesions,” in BioInformatics and BioEngineering, 2008. BIBE 2008. 8th IEEE International Conference on. IEEE, 2008, pp. 1–6. [Online]. Available: http://dx.doi.org/10.1109/BIBE.2008.4696799
[31] J. C. Bezdek, R. Ehrlich, and W. Full, “Fcm: The fuzzy c-means clustering algorithm,” Computers & Geosciences, vol. 10, no. 2, pp. 191–203, 1984. [Online]. Available: http://dx.doi.org/10.1016/0098-3004(84)90020-7
[32] D. S. Stoffer and K. D. Wall, “Bootstrapping state-space models: Gaussian maximum likelihood estimation and the kalman filter,” Journal of the american statistical association, vol. 86, no. 416, pp. 1024–1033, 1991. [Online]. Available: http://dx.doi.org/10.1080/01621459.1991.10475148
[33] M. Steinbach and P.-N. Tan, “knn: k-nearest neighbors,” The top ten algorithms in data mining, pp. 151–162, 2009. [Online]. Available: http://dx.doi.org/10.1201/9781420089653.ch8
[34] J. Alcon, C. Ciuhu, W. Kate, A. Heinrich, and N. Uzunbajakava, “Automatic imaging system with decision support for inspection of pigmented skin lesions and melanoma diagnosis,” IEEE Journal of Selected Topics in Signal Processing, vol. 3, no. 1, pp. 14–25, February 2009. [Online]. Available: http://dx.doi.org/10.1109/JSTSP.2008.2011156
[35] N. Otsu, “A threshold selection method from gray-level histograms,” Automatica, vol. 11, no. 285-296, pp. 23–27, 1975.
[36] M. E. Celebi, H. A. Kingravi, B. Uddin, H. Iyatomi, Y. A. Aslandogan, W. V. Stoecker, and R. H. Moss, “A methodological approach to the classification of dermoscopy images,” Computerized Medical Imaging and Graphics, vol. 31, no. 6, pp. 362–373, 2007.
[37] G. Capdehourat, A. Corez, A. Bazzano, R. Alonso, and P. Musé, “Toward a combined tool to assist dermatologists in melanoma detection from dermoscopic images of pigmented skin lesions,” Pattern Recognition Letters, vol. 32, no. 16, pp. 2187–2196, 2011. [Online]. Available: http://dx.doi.org/10.1016/j.patrec.2011.06.015
]]> [38] M. Fornaciali, S. Avila, M. Carvalho, and E. Valle, “Statistical learning approach for robust melanoma screening,” in Proceedings of the 2014 27th SIBGRAPI Conference on Graphics, Patterns and Images. IEEE Computer Society, 2014, pp. 319–326. [Online]. Available: http://dx.doi.org/10.1109/SIBGRAPI.2014.48[39] L. Y. Mera-González, J. A. Delgado-Atencio, J. C. Valdiviezo-Navarro, and M. Cunill-Rodríguez, “An algorithm for the characterization of digital images of pigmented lesions of human skin,” in SPIE Optical Engineering+ Applications. International Society for Optics and Photonics, 2014, pp. 921 718–921 718.
[40] Q. Abbas, M. Celebi, C. Serrano, I. F. García, and G. Ma, “Pattern classification of dermoscopy images: A perceptually uniform model,” Pattern Recognition, vol. 46, no. 1, pp. 86–97, 2013.
[41] R. Kaur, P. Albano, J. Cole, J. Hagerty, R. LeAnder, R. Moss, and W. Stoecker, “Real-time supervised detection of pink areas in dermoscopic images of melanoma: importance of color shades, texture and location,” Skin Research and Technology, 2015. [Online]. Available: http://dx.doi.org/10.1111/srt.12216
[42] C. Barata, M. Ruela, M. Francisco, T. Mendonça, and J. Marques, “Two systems for the detection of melanomas in dermoscopy images using texture and color features,” IEEE Syst. J, pp. 1–15, 2013. [Online]. Available: http://dx.doi.org/10.1109/JSYST.2013.2271540
[43] J. Scharcanski and M. E. Celebi, Computer vision techniques for the diagnosis of skin cancer. Springer, 2014. [Online]. Available: http://dx.doi.org/10.1007/978-3-642-39608-3
[44] A. Criminisi, P. Pérez, and K. Toyama, “Region filling and object removal by exemplar-based image inpainting,” Image Processing, IEEE Transactions on, vol. 13, no. 9, pp. 1200–1212, 2004. [Online]. Available: http://dx.doi.org/10.1109/TIP.2004.833105
[45] C. A. Cattaneo, L. I. Larchera, A. I. Ruggerib, A. C. Herreraa, and E. M. Biasoni, “Métodos de umbralización de imágenes digitales basados en entropia de shannon y otros,” Machine learning, vol. 20, no. 3, pp. 2785–2805, 2011.
[46] R. C. Gonzalez and R. E. Woods, “Digital imaging processing,” Massachusetts: Addison-Wesley, 1992.
[47] W. Torres, M. Landrove, M. Torreyes, and M. López, “Segmentación de imágenes dermatoscopicas en el espacio cielab utilizando filtros morfologicos SML,” in XII Congreso Internacional de Métodos Numéricos en Ingeniería y Ciencias Aplicadas, 2014.
]]> [48] Carrasco, Z., “Segmentación de Fallas en Soldaduras Utilizando Técnicas de Procesamiento Digital de Imágenes,” Universidad de Santiago de Chile, Facultad de Ingeniería, Master’s thesis, 2003.[49] M. Silveira, J. C. Nascimento, J. S. Marques, A. R. Marçal, T. Mendonça, S. Yamauchi, J. Maeda, and J. Rozeira, “Comparison of segmentation methods for melanoma diagnosis in dermoscopy images,” Selected Topics in Signal Processing, IEEE Journal of, vol. 3, no. 1, pp. 35–45, 2009. [Online]. Available: http://dx.doi.org/10.1109/JSTSP.2008.2011119