Services on Demand
Journal
Article
Related links
Share
Odontoestomatología
Print version ISSN 0797-0374On-line version ISSN 1688-9339
Odontoestomatología vol.21 no.33 Montevideo June 2019 Epub June 01, 2019
https://doi.org/10.22592/ode2019n33a2
Research
Comparative bacterial analysis of chronic and aggressive periodontitis in a sample population from Uruguay
1Cátedra Microbiología, Facultad de Odontología, Universidad de la República, Uruguay, andybad@hotmail.com
2Cátedra Periodoncia, Facultad de Odontología, Universidad de la República, Uruguay
3Cátedra Microbiología, Facultad de Odontología, Universidad de la República, Uruguay
Keywords: prevalence; microbiota; periodontitis
Objetivos:
analizar comparativamente la prevalencia de ciertos patógenos periodontales en cuadros de periodontitis agresiva y crónica según la clasificación vigente al momento de los estudios, en 101 pacientes uruguayos.
Métodos:
Este análisis se basa en los trabajos de detección de patógenos periodontales en pacientes con periodontitis crónica y agresiva en Uruguay mediante metodología convencional y molecular 1,2. Ambos trabajos analizaron los mismos microorganismos y utilizaron las mismas técnicas de recuperación y análisis. Se estudiaron Aggregatibacter actinomycetemcomitans, Porphyromonas gingivalis, Tannerella forsythia, Prevotella intermedia y Fusobacterium nucleatum.
Resultados:
En los cuadros crónicos destacó F. nucleatum, T. forsythia y P.gingivalis. En los agresivos F. nucleatum y P. intermedia, y dentro de éstos en los generalizados una mayor proporción de P. gingivalis y T. forsythia y de A. actinomycetemcomitans en los localizados.
Conclusiones:
Se pudo constatar que se mantienen los patrones documentados en la bibliografía en Uruguay.3,4
Palabras claves: prevalencia; microbiota; Periodontitis
Introduction and background
At the time of conducting and completing this work, the valid classification was that arising from the Workshop for a Classification of Periodontal Diseases and Conditions organized by the American Academy of Periodontology (AAP) in 1999 5-6.
According to this classification, chronic periodontitis is the most common type and typically progresses slowly. In contrast, aggressive periodontitis can lead to loss of tooth-supporting tissue in a short period. Aggressive periodontitis included localized aggressive periodontitis (LAP) and generalized aggressive periodontitis (GAP) according to the extent of periodontal destruction7.
Recently, a new classification of periodontal diseases based on stages, extent/distribution and grades has been agreed, without distinguishing between aggressive and chronic periodontitis. Staging is dependent on the severity and complexity of disease management (extent of damaged or lost tissue), the degree of extent and distribution compared to localized or more generalized cases, and grades mainly as an indication of rate of progression 8.
To correlate the clinical patterns of our studies with this new classification, regarding staging, we can say that our patients had a pocket depth > = 5 mm and a bone loss > = 2 mm.
As for extent and distribution, there is a difference regarding the distinction between generalized chronic and aggressive cases, and localized aggressive cases.
Finally, in grade there is a difference regarding the slower rate of progression of chronic cases in contrast to aggressive cases.
In periodontal disease, numerous studies based on bacterial specificity as responsible for the disease allowed us to focus on and establish the role of some microorganisms such as A. actinomycetemcomitans, P. gingivalis, T. denticola, P. intermedia and F. nucleatum in periodontal disease 9.
According to different studies, the microorganisms usually detected in chronic periodontitis belong to the red complex, such as P. gingivalis3,9-10.
In turn, the development of aggressive periodontitis has been linked to A. actinomycetemcomitans and P. gingivalis, the former being particularly important in localized cases and the latter in generalized cases 3,11-12. However, links have been found with another group of periodontopathogens such as T. forsythia, P. intermedia and F. nucleatum3,9.
In any case, these microorganisms do not act in isolation, but interact with each other, and this is how they determine the effects of disease 9.
Socransky et al. demonstrated that microorganisms in the subgingival plaque combine with each other creating bacterial complexes, and categorized them according to colors: green, yellow, blue, purple, orange, and red. These colors indicate the correlation of these complexes with the different degrees of severity of periodontal disease and its progression 11. The microorganisms that prevail in periodontal pathologies are the ones in Socransky’s red and orange complexes and also Aggregatibacter actinomycetemcomitans3,7,12-14.
The main differences between health and disease are based on the predominance of red and orange clusters. The red complex that appears later in biofilm development is formed by periodontopathogenic bacteria, namely: Porphyromonas gingivalis, Treponema denticola, Tannerella forsythia11,14.
On the other hand, A. actinomycetemcomitans plays a role in aggressive periodontitis, particularly in LAP, and it may also be important in the onset of the disease and then replaced by other obligate anaerobes as the disease progresses 12.
Up to the time of conducting these studies in patients in Uruguay, we knew that the relevance of the microorganisms studied in periodontal cases in the country was the same, but there were no local studies.
With the advent of molecular techniques, it was possible to determine the reality in our country without the risks of loss of viability implied when recovering samples in the laboratory using conventional techniques applied to periodontal microorganisms, mostly obligate anaerobes.
The microorganisms analyzed in this comparison were two members of the red complex, P. gingivalis and T. forsythia and two of the orange complex P. intermedia and F. nucleatum. It was interesting to include the latter in our studies given their ability to form bridges, which is relevant to biofilm formation 14.
We also studied A. actinomycetemcomitans, which has historically been linked to periodontitits, particularly LAP 3,12.
This study seeks to conduct a comparative analysis of the prevalence patterns of these five periodontal pathogens among the periodontitis cases studied.
Materials and methods
Samples were obtained from 101 selected patients with chronic and aggressive periodontitis from the Clinic of Periodontics of the School of Dentistry, Universidad de la República. Fifty-one patients with chronic periodontitis and 50 patients with aggressive periodontitis were studied; they had received no prior periodontal treatment and had signed the informed consent. Patients with diabetes, arthritis, ulcerative colitis, HIV, cancer and cardiovascular disease, pregnant women and those who had been treated with antibiotics and/or anti-inflammatories three months before the study were excluded. The Ethics Committee of the School of Dentistry, Universidad de la República, approved the design of both studies following MERCOSUR regulations and the Helsinki Declaration on research involving human subjects.
Supragingival plaque was taken to obtain the samples, using sterile gauze to avoid bleeding. Medium sterile paper points were placed deeply into the pocket and left for 15 seconds to collect the samples. They were then placed in 1.5 ml of RTF (Reduced Transport Fluid) 15. Each sample included eight medium sterile paper points (Nº 25) from four sites selected in each quadrant.
The samples were subjected to both traditional bacteriological techniques and molecular genetic techniques (PCR) at the Microbiology Department, School of Dentistry, Universidad de la República.
Traditional bacteriological diagnosis was made for the facultative Gram-negative bacillus A. actinomycetemcomitans and for P. gingivalis and P. intermedia (both pigmented strict anaerobic Gram-negative bacilli). We also conducted PCR identification for the microorganisms above and for T. forsythia and F. nucleatum. Samples were processed, stirred vigorously for 45 to 60 seconds, and then serial dilutions in RTF were prepared.
TSVB was used to recover A. actinomycetemcomitans: trypticase soy, bacitracin (75 µg/ml), vancomycin (5.0 µg/ml), 10 ml of horse serum (10%) through 1:10 dilution in RTF and incubation in a candle jar (to create capnophilic conditions) for 7 days at 37°C. Brucella Agar was used to recover the pigmented anaerobes: menadione (200 µg/ml) and hemin (5 mg/ml) and laked blood in 1:10, 1:100 and 1:1000 dilutions in RTF, streaking 100 µl of the last two dilutions and incubating in an anaerobic jar for 14 days at 37°C using the Oxoid® anaerobic indicator.
The original sample and 100 µl samples of the original sample were stored at -30°C for PCR processing.
The following strains were used as positive controls: Aggregatibacter actinomycetemcomitans (ATCC 29522), Porphyromonas gingivalis (BAA-308), Prevotella intermedia (ATCC 25611), Tannerella forsythia (ATCC 43037) and Fusobacterium nucleatum (ATCC 25586).
Identification of A. actinomycetemcomitans culture was performed by colony morphology, Gram staining, positive catalase test and negative MUG (4-Methylumbelliferyl-β-D-galactopyranoside) to study lactose fermentation, and also with bioMérieux® API kit in case of doubt.
Pigmented anaerobes were identified through pigment production, fluorescence to differentiate P. gingivalis from P. intermedia and bioMérieux® API kit in case of doubt 1-2.
The genome of each control strain was extracted using the BeadTMGenomic DNA Kit (as described by manufacturer Zymo Research®). DNA concentration (ng/µl) was quantified with NanoDrop( 2000.
Microbial genome databases were used to search for and select 16S rRNA sequences from the wild strain as targets in the design of species-specific and/or serotype-specific oligonucleotides. Additionally, we used bibliographic information from studies with validated oligonucleotides for the species 1-2.
Statistical analysis
Absolute frequencies in percentages were used. The populations were compared using the Student’s T-Test with a significance level of 0.05.
Results
As mentioned above, A. actinomycetemcomitans, P. gingivalis and P. intermedia were analyzed applying both traditional bacteriological and molecular techniques. We found a strong correlation in the results obtained through both techniques: a correlation equal to or higher than 85% (95% for A. actinomycetemcomitans, 93% for P. intermedia and 85% for P. gingivalis). When we found differences in the results obtained through the two techniques, the molecular technique result was considered valid and presented in this work.
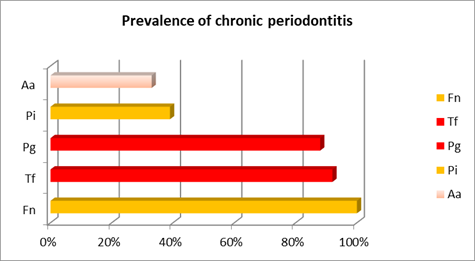
In chronic periodontitis, the most prevalent microorganism was F. nucleatum from the orange complex, occurring in 100% of patients. Additionally, most cases (approximately 90%) presented T. forshyta and P. gingivalis from the red complex. P. intermedia, from the orange complex, was recovered in 39% of patients, while A. actinomycetemcomitans was recovered in patients with chronic periodontitis but its prevalence was noticeably lower (33%) (Chart 1).
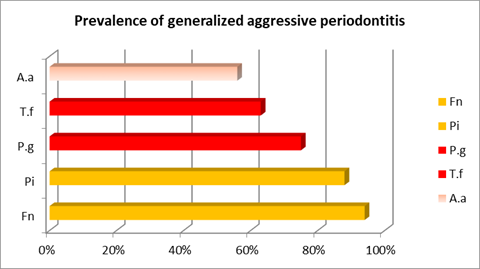
In both aggressive types of periodontitis, F. nucleatum, from the orange complex, was also important as it was the most prevalent microorganism in the generalized cases and in chronic periodontitis (Chart 2).
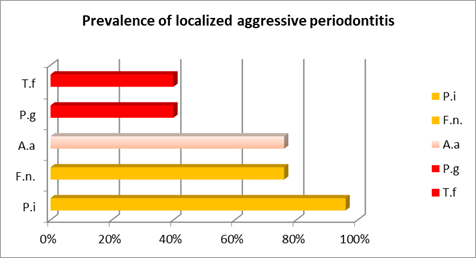
Prevotella intermedia, the other orange complex microorganism studied, was more prevalent than in the chronic cases, in particular in the localized cases, where it was the prevailing microorganism (Chart 3).
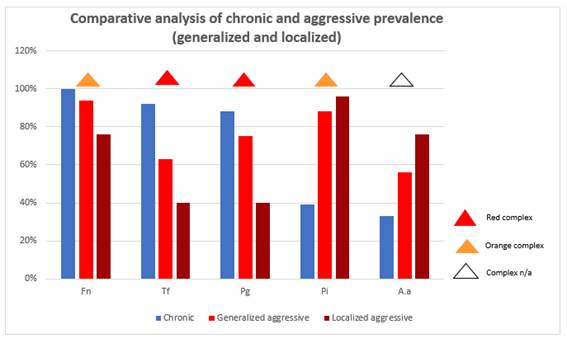
Furthermore, in aggressive cases, there was a higher prevalence of A. actinomycetemcomitans and a lower prevalence of the red complex microorganisms studied (P. gingivalis and T. forshytia) in relation to chronic cases. In aggressive cases, there are different results when distinguishing between generalized and localized cases. While the microorganisms from the red complex were more relevant in the generalized cases, A. actinomycetemcomitans prevailed in the localized cases, although in both cases there was higher prevalence than in chronic cases (Charts 2 and 3).
The following chart and table aim to facilitate the comparative analysis of the prevalence of each bacterial species studied in the different cases. (Chart 4 and Table 1)
Table 1: Comparative analysis of prevalence of chronic periodontitis, GAP and LAP
| Fn | Tf | Pg | Pi | Aa | |
|---|---|---|---|---|---|
| Chronic | 100% | 92% | 88% | 39% | 33% |
| GAP | 94% | 63% | 75% | 88% | 56% |
| LAP | 76% | 40% | 40% | 96% | 76% |
Fn= Fusobacterium nucleatum, Tf= Tannerella forsythia, Pg= Porphyromonas gingivalis, Pi= Prevotella intermedia, Aa = Aggregatibacter actinomycetemcomitans
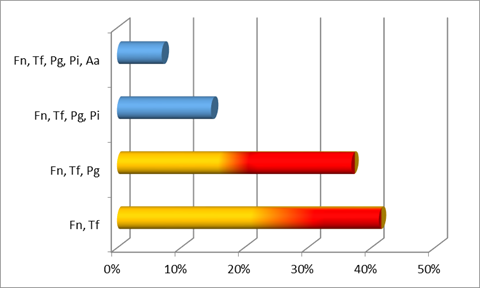
When considering the co-detection of species, where microorganisms are probably forming clusters or associations, we observed the following patterns for chronic periodontitis (Chart 5), generalized aggressive periodontitis (Chart 6) and localized aggressive periodontitis (Chart 7). As can be seen, the most common co-detection in the chronic cases was F. nucleatum with T. forsyhtia, followed by the association of F. nucleatum, T. forsythia and P. gingivalis (both red complex microorganisms).
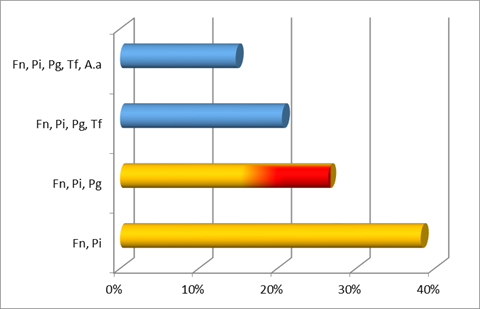
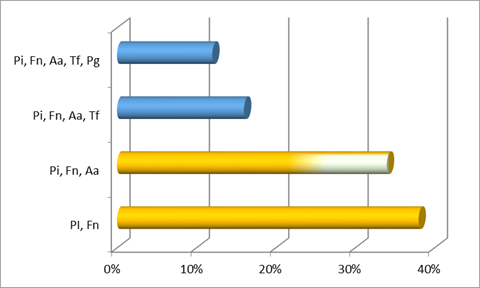
Additionally, the most common co-detection in the two aggressive types was F. nucleatum and P. intermedia. In generalized aggressive cases, the second main co-detection found was F. nucleatum, P. intermedia with P. gingivalis, while in localized aggressive cases it was F. nucleatum, P. intermedia (both from the orange complex) with A. actinomycetemcomitans.
The microbiota analyzed in the GAP cases was more similar to that of chronic periodontitis cases (greater representation of the red complex) than to that of LAP cases. Regarding A. actinomycetemcomitans and P. intermedia, it was more prevalent in both aggressive types than in chronic cases.
Discussion
Dental plaque is considered a biofilm. Microorganisms are not structured in a random order, which means that some species are more frequently associated with certain species than others 9,11.
Some species show a limited number of interactions with other species and others act like bridges, like F. nucleatum, which adheres to a vast number of species 16.
In the Uruguayan patients studied, the most prevalent microorganisms in chronic periodontitis were F. nucleatum from the orange complex, and T. forshyta and P. gingivalis from Socransky’s red complex.
It is not surprising for the last two microorganisms to be present in such high proportions in patients with chronic periodontitis, as they are microorganisms closely associated with the etiopathogenesis of the disease 3.
Furthermore, in the aggressive cases, these red complex microorganisms appeared in a lower rate than in chronic cases, particularly in localized cases. This pattern of higher prevalence of the red complex in the generalized versus the localized cases is consistent with studies conducted in other regions 13.
The high levels of F. nucleatum detected in both chronic and aggressive cases can be explained by its relevance in bacterial adhesion as it expresses multiple adhesins that allow late colonizers to bind. In addition, F. nucleatum may tolerate higher oxygen concentrations than P. gingivalis, which facilitates the generation of oxygen-reduction conditions. This promotes the growth and development of oxygen and other obligate anaerobes16. F. nucleatum can also raise pH through ammonia generation, which neutralizes the acid produced by the fermenting microorganisms, thus creating a more favorable environment for P. gingivalis and other acid-sensitive organisms 16.
P. intermedia, from the orange complex, had a higher prevalence in aggressive cases than in chronic cases. The fact that it appears at lower rates in chronic cases in our patients is similar to what other studies report, where subjects with high rates of P. gingivalis had little or no presence of P. intermedia3.
Our results, where P. intermedia was the most prevalent microorganism in aggressive cases, are comparable to those reported in other studies conducted in Germany and Africa 17-18.
In contrast, some studies in Korea and Australia report a greater presence of P. intermedia in chronic and generalized aggressive forms than localized aggressive ones 19-20.
We must remember that orange complex species such as P. intermedia and F. nucleatum allow for the colonization of red complex microorganisms 3, so the high numbers recovered for F. nucleatum in the three types of periodontitis and from both species in aggressive periodontitis (GAP and LAP) are revealing.
Regarding A. actinomycetemcomitans, there was a higher prevalence in aggressive cases versus chronic cases in our patients (and with greater emphasis on localized ones), which is compatible with the importance of this microorganism in aggressive cases, in particular in localized cases 3.
The studies conducted for the microorganisms analyzed and the comparison of the different periodontitis cases show that in Uruguay there is microbiota which is similar to that found in other regions 3,21. In aggressive cases, A. actinomycetemcomitans appeared in higher rates than in the generalized cases, which is compatible with the results of studies conducted in other countries 3,5.
Red-complex microorganisms T. forsythia and P. gingivalis, were particularly prominent in chronic processes, and even more in generalized aggressive cases than in localized ones.
Furthermore, F. nucleatum stood out in all cases, but particularly in chronic and generalized aggressive cases, where its role in the bacterial community is probably key. Something similar happened with P. intermedia from the orange complex, which prevailed in aggressive cases, particularly in localized cases.
We found a greater similarity in the microbiota studied between generalized aggressive cases and chronic cases than between localized aggressive cases and chronic cases. Additionally, aggressive cases have a higher rate of A. actinomycetemcomitans and P. intermedia compared to chronic cases, though they are more prevalent in LAP than in generalized cases
Referencias
1. Papone V, Verolo C, Zaffaroni L, Batlle A, Capo C, Bueno L, Gamonal J, Silva N, Soria S. Detección y prevalencia de patógenos periodontales de una población con periodontitis crónica en Uruguay mediante metodología convencional y metagenómica. Odontoestomatología. 2015; 17 (25): 23-32. [ Links ]
2. Badanian A, Ponce de León E, Rodríguez L, Bascuas T, Capo C, Batlle A, Bueno L, Papone V. Detección de patógenos periodontales de una población con Periodontitis Agresiva en Uruguay mediante metodología convencional y molecular. Odontoestomatología. 2018; 20 (32): 68-77. [ Links ]
3. Manakil J. Periodontal Diseases - A Clinician's Guide [en línea]. Rijeka: InTech; 2012. [Fecha de acceso: 1 de diciembre de 2018]. Disponible en: https://www.researchgate.net/publication/236935155_Periodontal_Diseases_A_Clinician%27s_Guide [ Links ]
4. Hinrichs E, Novak MJ. Clasificación de las enfermedades y condiciones que afectan el periodonto. En: Newman MG, Takei H, Klokkevold PR, Carranza FA. Periodontología Clínica de Carranza. 11a ed. Venezuela: Amolca, 2014. p 60-67 [ Links ]
5. Novak KF, Novak MJ. Periodontitis agresiva. En: Newman M.G., Takei H, Klokkevold PR, Carranza FA. Periodontología Clínica de Carranza. 11a ed. Venezuela: Amolca, 2014. p 232-236. [ Links ]
6. Armitage GC. Development of a Classification System for Periodontal Diseases and Conditions. Ann Periodontol. 1999; 4 (1): 1-6. [ Links ]
7. Bascones Martínez A, Figuero Ruiz E. Periodontal diseases as bacterial infection. Av Periodon Implantol. 2005; 17 (3): 111-118. [ Links ]
8. Caton JG, Armitage G, Berglundh T, Chapple ILC, Jepsen S, Kornman KS, Mealey BL, Papapanou PN, Sanz M, Tonetti MS. A new classi?cation scheme for periodontal and peri-implant diseases and conditions - Introduction and key changes from the 1999 classi?cation. J Periodontol. 2018; 89 (1): 1-8. [ Links ]
9. Feres M, Cortelli SC, Figueiredo LC, Haffajee AD, Socransk SS. Microbiological Basis for periodontal therapy. J Appl Oral Sci. 2004; 12 (4): 256-266. [ Links ]
10. Farias BC, Souza PRE, Ferreira B, Melo RSA, Machado FB, Gusmão ES, Cimões R. Occurrence of periodontal pathogens among patients with chronic periodontitis. Braz. J. Microbiol. 2012; 43(3): 909-916 [ Links ]
11. Socransky SS, Haffajee AD. Periodontal microbial ecology. Periodontol 2000, 2005; 38: 135-187. [ Links ]
12. Nibali L. Aggressive Periodontitis: microbes and host response, who to blame?. Virulence. 2015; 6 (3): 223-228. [ Links ]
13. Könönen E, Müller HP. Microbiology of aggressive periodontitis. Periodontol 2000. 2014; 65 (1): 46-78. [ Links ]
14. Holt SC, Ebersole JL. Porphyromonas gingivalis, Treponema denticola, and Tannerella forsythia: the 'red complex', a prototype polybacterial pathogenic consortium in periodontitis. Periodontol 2000. 2005; 38: 72-122. [ Links ]
15. Syed SA, Loesche WJ. Survival of Human Dental Plaque Flora in Various Transport Media. App. Microbiology. 1972; 24 (4): 638-644. [ Links ]
16. Wright CJ, Burns LH, Jack AA, Back CR, Dutton LC, Nobbs AH, Lamont RJ, Jenkinson HF. Microbial interactions in building of communities. Mol Oral Microbiol. 2013; 28 (2): 83-101. [ Links ]
17. Nonnenmacher C, Mutters R, Flores de Jacoby L. Microbiological characteristics of subgingival microbiota in adult periodontitis, localized juvenile periodontitis and rapidly progressive periodontitis subjects. Clin Microbiol Infect. 2001; 7 (4): 213-217. [ Links ]
18. Amel Y, Bouziane D, Leila M, Ahmed B. Microbiological Study of Periodontitis in the West of Algeria. World J. Med. Sci. [En línea] 2010; 5(1): 7-12 [ Links ]
19. Lee JW, Choi BK, Yoo YJ, Choi SH, Cho KS, Chai JK, Kim CK. Distribution of periodontal pathogens in Korean aggressive periodontitis. J Periodontol. 2003; 74 (9): 1329-1335. [ Links ]
20. Oettinger-Barak O, Sela MN, Sprecher H, Machtei EE. Clinical and microbiological characterization of localized aggressive periodontitis: a cohort study. Aust Dent J. 2014; 59: 165-171. [ Links ]
21. Ximenez-Fyvie LA, Almaguer-Flores A, Jacobo-Soto V, Lara-Cordoba M, Moreno- Borjas J-Y, Alcantara-Maruri E. Subgingival microbiota of periodontally untreated Mexican subjects with generalized aggressive periodontitis. J Clin Periodontol. 2006; 33: 869-877. [ Links ]
Received: February 18, 2019; Accepted: April 26, 2019










 text in
text in