Serviços Personalizados
Journal
Artigo
Links relacionados
Compartilhar
Odontoestomatología
versão impressa ISSN 0797-0374versão On-line ISSN 1688-9339
Odontoestomatología vol.20 no.32 Montevideo dez. 2018
https://doi.org/10.22592/ode2018n32a9
Research
Detection of periodontal pathogens in a Uruguayan population with aggressive periodontitis using conventional and molecular methods
1 Cátedra de Microbiología, Facultad de Odontología. Universidad de la República, Uruguay
2 Cátedra de Microbiología, Facultad de Odontología, Universidad de la República, Uruguay
3 Ciencias Biológicas, Facultad de Ciencias, Universidad de la República, Uruguay
4 Ciencias Biológicas, Facultad de Ciencias, Universidad de la República, Uruguay
5 Cátedra de Periodoncia, Facultad de Odontología, Universidad de la República, Uruguay
6 Cátedra de Periodoncia, Facultad de Odontología, Universidad de la República, Uruguay
7 Cátedra de Periodoncia, Facultad de Odontología, Universidad de la República, Uruguay
8 Cátedra de Microbiología, Facultad de Odontología, Universidad de la República, Uruguay. vpaponey@gmail.com
Keywords: prevalence; microbiota; aggressive periodontitis
Palabras clave: prevalencia; microbiota; Periodontitis Agresiva
Introduction and background
Aggressive periodontitis is a destructive periodontal disease that can affect the health of the primary or permanent dentition and, over a short period, can lead to loss of tooth-supporting tissue. The term aggressive periodontitis was approved at the Workshop for the Classification of the Periodontal Diseases and Conditions organized by the American Academy of Periodontology (AAP) in 1999 to define a group of destructive periodontal diseases with a rapid progression rate, a classification that was valid at the time this study was conducted 2-4.
Aggressive periodontitis can appear in its localized form, localized aggressive periodontitis (LAP) or in its generalized form, generalized aggressive periodontitis (GAP) depending on the extent of periodontal destruction 1.
According to the World Health Organization (WHO), severe periodontal disease, which may result in tooth loss, affects 15-20% of middle-aged (35-44 years) adults 5. As for the aggressive form, Albandar mentions that the prevalence of aggressive periodontitis is variable if different locations are considered, although many of these studies used different variables and methodologies. The prevalence of localized aggressive periodontitis varies considerably between continents, and differences in race/ethnicity seem to be a major contributing factor. Several studies have consistently shown that aggressive periodontitis is more prevalent in Africa and Afrodescendants and less prevalent in the Caucasian population of Europe and North America. The prevalence among children and young adults is higher in older than in younger age groups 6. The prevalence of aggressive periodontitis in Europe is low, between 0.1% and 0.2%, and prevalence is higher, between 3% and 10%, in Brazil and the United States 7. Studies show that in Latin America the prevalence of aggressive periodontitis is higher in this region than in industrialized countries. Prevalence ranges from 0.3% to 4.5%, and the localized form is the least prevalent one 8. Worldwide, the prevalence of LAP is lower than 1%, and the prevalence of GAP is 0.13% 9-10.
Aggressive periodontitis is a disorder of multifactorial origin, such as host response, genetic factors and the subgingival microbiota 6,9,11.
Some of the bacterial pathogenic species that have been linked to the development of aggressive periodontitis are: Aggregatibacter actinomycetemcomitans (Aa)2,12 and Porphyromonas gingivalis (Pg) ( 13, the first being particularly important in the localized form 14-15 and the second one in the generalized form 15-16, but another group of periodontopathogens, including Tannerella forsythia (Tf), Prevotella intermedia (Pi)1-15 and Fusobacterium nucleatum (Fn)15) have been linked to it.
Studies have been conducted in Uruguay using traditional bacterial identification methods, but so far, there are no studies concerning the levels of simultaneous occurrence of the periodontal pathogens mentioned above in cases of aggressive periodontitis. In addition, traditional bacteriological techniques have certain limitations, such as maintaining the viability of the sample, bacterial growth time (7 to 15 days), as well as the difficulty of bacterial growth since they respond to microorganisms that require specific nutrients and are, mostly, obligate anaerobes. Bacterial recognition involves performing additional tests, such as biochemical tests, to identify the species 17.
The introduction of molecular analysis methods, such as polymerase chain reaction (PCR), has expanded and simplified studies of the composition of oral microbiota of patients with periodontal pathologies, improving their sensitivity, specificity and efficiency 18.
In this work, we analyzed the presence of five periodontal pathogens: Aggregatibacter actinomycetemcomitans, Porphyromonas gingivalis, Prevotella intermedia, Tannerella forsythia and Fusobacterium nucleatum in the periodontal pockets of Uruguayan patients from the Clinic of Periodontics of the School of Dentistry, Universidad de la República, Uruguay. Microbiological identification techniques included traditional bacteriological techniques, as well as techniques to identify the genetic material present through PCR 17.
In this study, we were able to identify the microorganisms that are present and more prevalent in cases of aggressive periodontitis in Uruguay, both in the generalized and localized forms, and to determine whether the prevalence rates are the same as those found in other geographical locations.
It also allowed us to correlate traditional bacteriological techniques and molecular techniques.
Work methodology
We selected 50 patients from the Clinic of Periodontics of the School of Dentistry with aggressive periodontitis, without prior periodontal treatment and who signed an informed consent. Patients with diabetes, arthritis, ulcerative colitis, HIV, cancer and cardiovascular disease, pregnant women and those who had been treated with antibiotics and/or anti-inflammatories three months before the study were excluded. The Ethics Committee of the School of Dentistry, UdelaR, approved the design of the study following the regulations of the MERCOSUR and the Helsinki Declaration on research involving human subjects.
Supragingival plaque was taken to obtain the samples, using sterile gauze to avoid bleeding. Medium sterile paper points (Nº 25) were placed deeply into the pocket and left for 15 seconds to collect the samples. They were then placed in 1.5 ml of RTF (Reduced Transport Fluid) 18. Each sample included eight paper points from four sites selected in each quadrant.
We carried out the bacteriological diagnosis using the traditional methodology for A. actinomycetemcomitans and for the pigmented anaerobes P. gingivalis and P. intermedia. We also conducted the identification by PCR for T. forsythia and F. nucleatum. Samples were processed in the Microbiology Laboratory of the School of Dentistry, stirred vigorously for 45 to 60 seconds, and then serial dilutions in RTF were prepared. 100 µl was taken from the sample and placed in an Eppendorf tube with 900 µl more of RTF, thus achieving a 1:10 dilution. 100 µl was streaked, expanding the sample with a glass rod on a TSVB plate: tryptic soy serum, bacitracin (75 µg/ml), vancomycin (5.0 µg/ml), 10 ml of horse serum (10%) to isolate A. actinomycetemcomitans. The plates were incubated in a candle jar (environment rich in carbon dioxide) for 7 days at 37ºC. In addition, 100 µl was taken from the RTF original sample and 1:10, 1:100 and 1:1000 dilutions in RTF were obtained. Then, 100 µl of the latter two was streaked in base agar medium with blood with menadione and hemin, and incubated for 14 days at 37ºC in absolute anaerobiosis to recover anaerobes (Porphyromonas and Prevotella).
From the sample in RTF, 100 µl samples were taken, in addition to the original sample, stored at -30ºC and then processed using the PCR technique. The following wild strains were used as positive controls: Aggregatibacter actinomycetemcomitans (ATCC 29522), Porphyromonas gingivalis (BAA-308), Prevotella intermedia (ATCC 25611), Tannerella forsythia (ATCC 43037) and Fusobacterium nucleatum (ATCC 25586).
Identification by culture
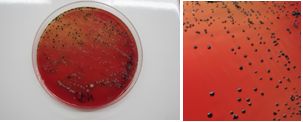
Presumptive colonies of A. actinomycetemcomitans were identified given the presence of a star-like structure inside the colonies observed using a stereoscopic magnifying glass (Fig. 1), Gram stain technique, catalase test (+) and MUG negative (4-Methylumbelliferyl-β-D-galactoside to study lactose fermentation).
Pigmented anaerobic bacteria (Porphyromonas and Prevotella) were identified by pigment production (Fig. 2). They were also observed under ultraviolet light (360 nm), for differentiation based on fluorescence, in which colonies of P. gingivalis do not fluoresce and colonies of P. intermedia do (Fig. 3). To avoid any doubt, the identification was performed with biochemical tests for anaerobes using the API® kit.
Fluorescent colonies of Prevotella intermedia under ultraviolet light
A control group of 10 selected patients who did not have periodontitis was created to validate the study. As a result, no growth was obtained in 8 out of the 10 cases in the selective medium for A. actinomycetemcomitans. The remaining two did show growth, but it was not A. actinomycetemcomitans. As for pigmented anaerobes, growth was obtained in 4 out of the 10 cases, but with a low count.
Identification by PCR. Extraction of microbial genomic DNA
The samples were defrosted and homogenized for 30 seconds rigorously with a vortex. The content was then transferred to another Eppendorf tube and centrifuged at 13500 rpm for 3 minutes. The supernatant was discarded and the pellet was suspended again by pipetting in 500 µl of RTF for 5 minutes at 100ºC. They were then placed in ice for 5 minutes and then centrifuged at 13500 rpm for 5 minutes at 4ºC. Finally, the supernatant with the microbial genomic DNA was taken and stored at -30ºC until the PCR was performed.
The genome of each wild control strain was extracted using the ZymoBeadTM Genomic DNA Kit (as described by the manufacturer). DNA concentration (ng/µl) was quantified by NanoDrop 2000.
The NCBI (National Center of Biotechnology Information) database was used to design the primers in order to search for and select 16S rRNA sequences to be targeted by the specific primers for T. forsythia (GenBank: AP013045.1) and F. nucleatum (GenBank: MH078456.1). Bibliographic information of studies with validated oligonucleotides for the species was also used 19.
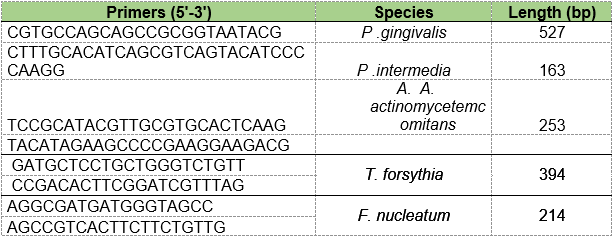
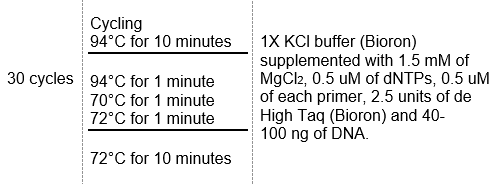
PCR was performed with each pair of oligonucleotides using the genomes extracted from the wild strain. The primers used to detect bacterial species A. actinomycetemcomitans, P. gingivalis, P. intermedia, T. forsythia and F. nucleatum are shown in Table 1. The conditions of the multiplex PCR used for detection of these microorganisms are shown in Tables 2 and 3.
Table 3: Conditions of the multiplex PCR for detection of P. gingivalis, P. intermedia, A. actinomycetemcomitans
The results of amplifications (bands) were viewed in 1.2% agarose gel, 0.5X TBE (Tris-borate-EDTA) buffer, 5 µl of GoodView, 2 µl of Cyan/Orange loading buffer; 2-5 µl of the PCR product for 2 h at 80 V. They were also photographed using a digital camera placed over an ultraviolet (UV) light transilluminator (Figs. 4 and 5).
Statistical analysis
Descriptive statistics were applied using absolute frequencies in percentages. The populations were compared using the Student’s T-Test with a significance level of 0.05.
Results
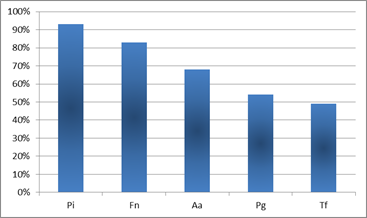
The most prevalent microorganisms in the cases of aggressive periodontitis were P. intermedia and F. nucleatum. They were present in 93% and 83% of the patients, respectively. The next one in incidence was A. actinomycetemcomitans, a microorganism that is strongly correlated with aggressive periodontitis 9,11-12,16.
A. actinomycetemcomitans appeared in around 70% of the patients studied (Chart 1).
When distinguishing between localized and generalized aggressive periodontitis, the microbiota is differentiated by emphasizing the different mirobiological etiopathogenesis of both forms. Of all the cases of aggressive disease studied, 60% were localized cases and 40% generalized cases. This is why in an analysis of the microbiota of aggressive cases of the disease without distinguishing between localized and generalized forms, prevalence percentages are conditioned by the larger number of localized cases.
Generalized aggressive periodontitis
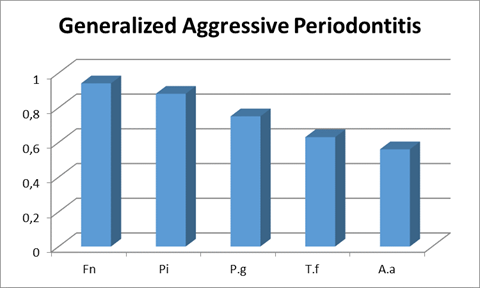
Although F. nucleatum was a microorganism with a high prevalence in both forms, it mainly stood out in the cases of generalized aggressive disease, as it was present in almost 95% of the patients.
In the cases of the generalized form, the next microorganism in prevalence was P. intermedia (around 90%), followed by P. gingivalis in 75% of the samples and by T. forsythia in 63% of them. Moreover, although A. actinomycetemcomitans was recovered in several patients, it was the least prevalent of the microorganisms studied, being present in about 56% of cases (Chart 2).
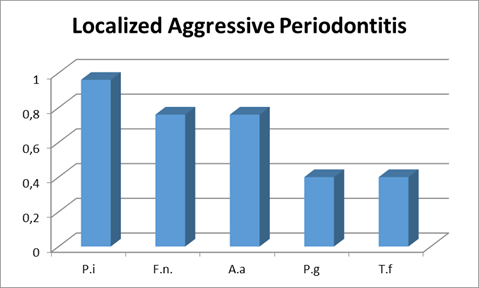
Localized aggressive periodontitis
As for the localized form, the most prevalent microorganism was P. intermedia, which was present in 96% of patients, followed by F. nucleatum and A. actinomycetemcomitans, both with a prevalence of around 75%. It is worth noting that there was a higher prevalence of A. actinomycetemcomitans than in the generalized form of the disease. This is consistent with the relevance that this microorganism has in localized cases of the disease 13-15. Finally, the microorganisms P. gingivalis and T. forsythia had the lowest prevalence, being found in 40% of the patients studied (Chart 3).
Comparison of microbial prevalence between GAP and LAP
To facilitate the comparison, the respective prevalences of the different pathogens in the generalized and localized forms in our studies with Uruguayan patients are presented in a consolidated form below (Chart 4).
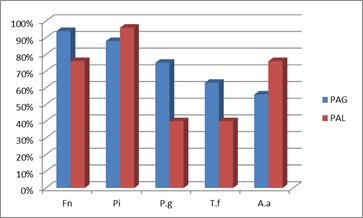
Chart 4: Comparison of the prevalence of periodontal pathogens between generalized (GAP) and localized aggressive periodontitis (LAP)
Proportion of pathogens studied in the cases of aggressive periodontitis
Another way to analyze the relevance of the microorganism in the disease is to study it in connection to the detection of the rest of the pathogens studied. If we consider the detection of a certain pathogen in a patient as one occurrence, 100% of the occurrences would be represented by the total number of detections of the five pathogens studied. The following proportions can be found by studying the proportion of each microorganism in relation to the rest of the microorganisms (the total number of occurrences) (Chart 5).
The most prevalent microorganism was P. intermedia, followed by F. nucleatum. A. actinomycetemcomitans was in third place with 20%.
As mentioned above, percentages are largely influenced by the larger number of localized over generalized cases.
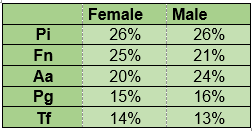
No significant differences were found in the proportion of microorganisms when taking into account sex with a 0.05 significance level using the t-test for two samples, assuming equal variances (Table 4).
Discussion
The microorganisms present in generalized aggressive periodontitis (GAP) in our study are similar to those found in cases of aggressive periodontitis in other countries 15-16. According to the literature, the microorganisms usually detected in cases of GAP are P. gingivalis, A. actinomycetemcomitans and T. forsythia16.
P. gingivalis is especially more prominent than A. actinomycetemcomitans in cases of generalized disease 15.
As for the presence of the pathogens analyzed in our study, we found that in generalized aggressive periodontitis, P. gingivalis was more common than A. actinomycetemcomitans, which is consistent with the literature and other locations such as Chile and Colombia 20-21.
As for localized aggressive periodontitis, according to our results, A. actinomycetemcomitans exceed the periodontopathogens P. gingivalis and T. forsythia, similarly to other locations such as Europa and North America 2,11,15,21, where A. actinomycetemcomitans is recovered particularly in cases of localized disease. However, in studies carried out in Latin America (Chile, Colombia), the recovery of P. gingivalis and T. forsythia exceeded that of A. actinomycetemcomitans in this form of the disease 20,22-23.
Furthermore, some authors highlight the importance of A. actinomycetemcomitans given its role in the pathogenesis of aggressive periodontitis, especially in the localized form, even though it has been suggested that it is probably of particular relevance at the beginning of the pathology and is then replaced by other pathogenic bacteria as the disease progresses 11,13.
Conclusions
It was found that there is a difference in the flora between the localized and generalized forms of the disease in the Uruguayan patients studied.
The most prevalent microorganism in localized aggressive periodontitis was P. intermedia, while in generalized aggressive periodontitis it was F. nucleatum.
In addition, P. gingivalis and T. forsythia are more prevalent in the generalized form than in the localized form of the disease.
We also confirmed the significance of A. actinomycetemcomitans in aggressive periodontitis, which is especially significant in localized cases.
A similar flora to other geographical locations was present in aggressive cases of periodontitis in our country, even though in some of those P. gingivalis is more common than A. actinomycetemcomitans in all forms of aggressive periodontitis (20, 22). In our case, we only found this relation in generalized cases of the disease. In addition, A. actinomycetemcomitans was present in a higher proportion than P. gingivalis in cases of the localized form of the disease, which is consistent with the data reported in the literature 16.
It was also possible to correlate the traditional bacteriological study with molecular study techniques
Referencias
1. Bascones-Martínez A, Figuero Ruiz E. Periodontal diseases as bacterial infection. Av Periodon Implantol. 2005; 17 (3): 111-118. [ Links ]
2. Hinrichs E, Novak MJ. Clasificación de las enfermedades y condiciones que afectan el periodonto. En: Newman MG, Takei H, Klokkevold PR, Carranza FA. Periodontología Clínica de Carranza. 11a ed. Venezuela: Amolca, 2014. p 60-67 [ Links ]
3. Armitage, GC. Development of a Classification System for Periodontal Diseases and Conditions. Ann Periodontol. 1999; 4 (1): 1-6. [ Links ]
4. G Caton J, Armitage G, Berglundh T, Chapple ILC, Jepsen S, S Kornman K, L Mealey B, Papapanou PN, Sanz M, S Tonetti M. A new classification scheme for periodontal and peri-implant diseases and conditions - Introduction and key changes from the 1999 classification. J Clin Periodontol. 2018; 45 (20): s1-8. [ Links ]
5. OMS. Salud bucodental Nota informativa No 318 Abril 2012, Fecha de acceso: 26 de noviembre de 2017 Disponible en: http://www.who.int/mediacentre/factsheets/fs318/es/ [ Links ]
6. Albandar JM. Aggressive and acute periodontal diseases. Periodontol 2000. 2014; 65: 7-12. [ Links ]
7. Chan-Myung C, Hyung-Keun Y, Seong-Nyum J. The clinical assessment of aggressive periodontitis patients. J Periodontal Implant Sci. en línea 2011, 41(3): 143-148. Fecha de acceso: 17 de diciembre de 2017 . Disponible en: http://europepmc.org/articles/PMC3139048 [ Links ]
8. Oppermann, RV. An overview of the epidemiology of periodontal diseases in Latin America. Braz Oral Res en línea 2007, 21: 8-15. Fecha de acceso: 10 de diciembre de 2017 . Disponible en: https://dx.doi.org/10.1590/S1806-83242007000500003 [ Links ]
9. Joshipura V, Yadalam U, Brahmavar B. Aggressive periodontitis: A review. J Int Clin Dent Res Organ. 2015; 7 (1): 11-17. [ Links ]
10. Sharmal P, Vishnoi S, Chandran S, Rathod D. Generalized Aggressive Periodontitis, Multifactorial treatement modalities - A case report, IJAR en línea 2016, 4 (5): 1053-1057. Fecha de acceso: 10 de diciembre de 2017 . Disponible en: http://www.journalijar.com/uploads/532_IJAR-10305.pdf [ Links ]
11. Könönen E , Müller HP. Microbiology of aggressive periodontitis. Periodontol 2000. 2014; 65 (1): 46-78. [ Links ]
12. Ardila Medina CM, Botero Zuluaga L, Guzmán Zuluaga IC. Comparación de las características sociodemográficas, clínicas y microbiológicas de pacientes con periodontitis agresiva y crónica. AMC En línea 2014; 18 (5): 532-544. Fecha de acceso: 10 de diciembre de 2017 . Disponible en: http://scielo.sld.cu/scielo.php?script=sci_arttext&pid=S1025-02552014000500009 [ Links ]
13. Nibali L. Aggressive Periodontitis: microbes and host response, who to blame?. Virulence En línea 2015; 6 (3): 223-228. Fecha de acceso: 10 de diciembre de 2017 . Disponible en: http://europepmc.org/articles/PMC4601283 [ Links ]
14. Avila-Campos MJ, Simionato MRL, Cai S, Mayer MPA, De Lorenzo JL, Zelante F. Virulence factors of Actinobacillus actinomycetemcomitans: other putative factors. Pesqui. Odontol. Bras. 2000; 14 (1): 05-11. [ Links ]
15. Manakil J. Periodontal Diseases - A Clinician's Guide en línea . Rijeka: InTech; 2012. Fecha de acceso: 22 de noviembre de 2017 . Disponible en: https://www.researchgate.net/publication/236935155_Periodontal_Diseases_-_A_Clinician%27s_Guide [ Links ]
16. Novak KF, Novak MJ. Periodontitis agresiva. En: Newman MG., Takei H, Klokkevold PR, Carranza FA. Periodontología Clínica de Carranza. 11a ed. Venezuela: Amolca, 2014. p 232-236 [ Links ]
17. Frías López MC, Uria Ovando V, Carasol Campillo M. Métodos de diagnóstico microbiológico en la enfermedad periodontal. Cient. Dent En línea 2009; 6 (2): 93-101. Fecha de acceso: 9 de diciembre de 2017 . Disponible en http://www.coem.org.es/sites/default/files/revista/cientifica/vol6-n2/17-25.pdf [ Links ]
18. Syed SA, Loesche WJ. Survival of Human Dental Plaque Flora in Various Transport Media. App. Microbiol. 1972; 24 (4): 638-644. [ Links ]
19. García L, Tercero JC, Legido B, Ramos JA, Alemany J, Sanz M. Rapid detection of Actinobacillus actinomycetemcomitans, Prevotella intermedia and Porphyromonas gingivalis by multiplex PCR. J Periodontal Res. 1998; 33 (1): 59-64. [ Links ]
20. Gajardo M, Silva N, Gómez L, León R, Parra B, Contreras A, Gamonal J. Prevalence of Periodontophatic Bacteria in Aggressive Periodontitis Patients in a Chilean Population. J Periodontol. 2005; 76: 289-294. [ Links ]
21. Mayorga-Fayad I, Lafaurie GI, Contreras A, Castillo DM, Barón A, Aya MR. Microflora subgingival en periodontitis crónica y agresiva en Bogotá, Colombia: un acercamiento epidemiológico. Biomédica. 2007; 27 (1): 21-33. [ Links ]
22. Botero JE, Contreras A, Lafaurie G, Jaramillo A, Betancourt M, Arce RM. Occurrence of Periodontopathic and Superinfecting Bacteria in Chronic and Aggressive Periodontitis Subjects in a Colombian Population. J Periodontol. 2007; 78 (4): 696-704. [ Links ]
23. Contreras A, Moreno SM, Jaramillo A, Pelaez M, Duque A, Botero JE, Slots J. Periodontal microbiology in Latin America. Periodontol 2000. 2015; 67: 58-86 [ Links ]
Received: February 14, 2018; Accepted: August 07, 2018










 texto em
texto em